This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
How an ultra-sensitive on-off switch helps axolotls regrow limbs
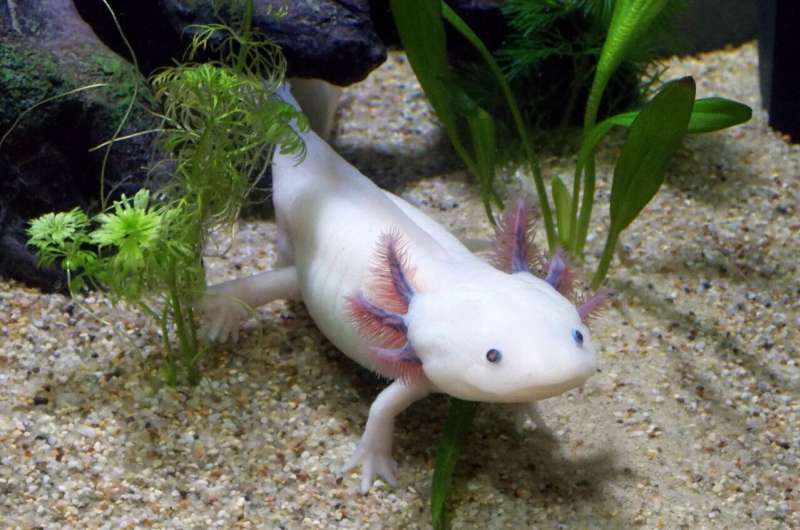

It's one of the mysteries of nature: How does the axolotl, a small salamander, boast a superhero-like ability to regrow nearly any part of its body? For years, scientists have studied the amazing regenerative properties of the axolotl to inform wound healing in humans.
Now, Stanford Medicine researchers have made a leap forward in understanding what sets the axolotl apart from other animals. Axolotls, they discovered, have an ultra-sensitive version of mTOR, a molecule that acts as an on-off switch for protein production. And, like survivalists who fill their basements with non-perishable food for hard times, axolotl cells stockpile messenger RNA molecules, which contain genetic instructions for producing proteins.
The combination of an easily activated mTOR molecule and a repository of ready-to-use mRNAs means that after an injury, axolotl cells can quickly produce the proteins needed for tissue regeneration. The new findings were published July 26 in Nature.
"Until now, it has been difficult to pinpoint a specific change in a single molecule in axolotls that was so critical for regenerative potential," said Maria Barna, an associate professor of genetics and the senior author of the paper. "We've made significant headway toward understanding how we may eventually manipulate the mTOR pathway to boost regenerative potential in humans."
From mRNA to protein
In the past, researchers trying to figure out how the axolotl regrows entire body parts—including legs, tails, eyes and even the heart—focused on how levels of mRNA molecules changed after an axolotl has an injury. Scientists have long used mRNA molecule levels as a proxy for protein levels; after all, mRNA must exist before a protein can be produced. However, these studies only shed light on what happens to the production of mRNA molecules after injury—not what happens to the translation of mRNA into protein products.
"There are hundreds of mRNA transcripts that appear after a wound, but researchers were really struggling to figure out what it was about salamanders that could explain their regenerative potential," Barna said.
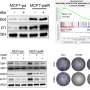
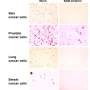
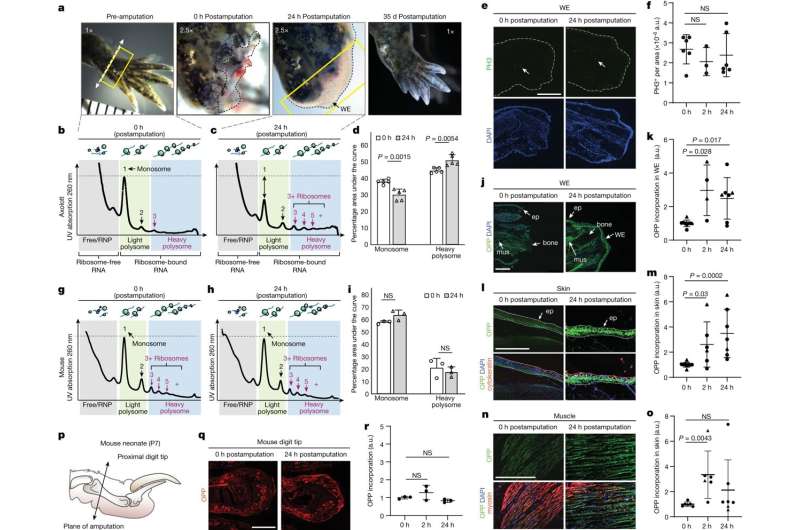
Her lab took a different approach, focusing on which mRNA molecules near a wound were attached to ribosomes, little molecular machines that create proteins. That helped the scientists zero in on which proteins were being made, rather than which mRNA molecules loitered near the injury site. Usually, when cells encounter stress (such as after an injury) they decrease overall protein production to save energy, so Barna's group expected to see fewer mRNA molecules bound to ribosomes. Instead, they saw more.
"It was a 180-degree flip when we realized that when an axolotl loses a limb, it actually increases protein synthesis despite the energy cost," Barna said.
Further experiments showed that axolotl cells "stockpile" mRNA, translating less than 20% of it at any given time. When the researchers analyzed how axolotls respond to injury, they found that protein synthesis is activated, leading to the translation of hundreds of stockpiled transcripts. That long-term storage also explained the speed at which protein synthesis occurred during regeneration.
"We had a gut feeling that looking at protein synthesis more closely would be important, " said Olena Zhulyn, Ph.D., postdoctoral scholar and lead author of the study. "But never in a million years did we expect that protein synthesis would be the key to the mystery of the axolotl's regeneration."
A connection to mTOR
A question remained: What was activating the mRNAs and causing them to bind to ribosomes after axolotls lose a body part? The researchers noticed that many of the stockpiled mRNA molecules had a shared sequence of nucleotides at one end of the mRNA which was known to be regulated by the enzyme mTOR to promote protein production.
The research found that the axolotl mTOR protein is highly sensitive—the axolotl variety contained a genetic alteration, an expansion in sequence, seen only in axolotl and related salamanders.
Investigating further, Barna and her team collaborated with researchers at University of California, San Francisco to probe the structural differences between axolotl mTOR and mammalian mTOR.
In humans and mice, mTOR (and resulting protein production) activates only when there's a surplus of nutrients. In other words, mammalian cells use mTOR to make proteins only in the best of times. But in axolotls, after an injury causes cell damage and the breakdown of many molecules, the small rush in loose nutrients is enough to flip the ultra-sensitive mTOR to its active state, turning on the cellular factories that make new proteins.
"Finding this genetic change was a shock—mTOR is an ancient enzyme that is the same in virtually all organisms," said Zhulyn. "But in axolotls we were seeing evolution of new sequences and a structure that changed its fundamental properties."
When Barna and her colleagues blocked mTOR with a drug used to prevent protein production and cell division in cancers, the animals were no longer able to regrow limbs. The axolotl mTOR is hypersensitive to stimulation (in this case, injury) but is not more active than mammalian mTOR, they found. That's key, said Barna—hyperactive mTOR has been linked to tumor growth in many human cancers. Given that the axolotl mTOR doesn't show hyperactivity, that could explain the remarkable cancer resistance seen in axolotls, she said.
More research is needed to probe whether changing or stimulating mTOR in humans could improve wound healing or spur the regeneration of damaged, diseased organs, Barna said.
"I think there are a still a lot of lessons to be learned about how this tight control of mRNA translation is allowing wound healing and tissue regeneration," said Barna. "There is a whole new world to be discovered when it comes to both the basic biology of translation and healing."
More information: Olena Zhulyn et al, Evolutionarily divergent mTOR remodels translatome for tissue regeneration, Nature (2023). DOI: 10.1038/s41586-023-06365-1
Salamanders' regenerative potential might be driven by a specific protein variant, Nature (2023). DOI: 10.1038/d41586-023-02111-9
Journal information: Nature
Provided by Stanford University