This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Mapping the evolution of E. coli's main virulence factor offers a refined drug target
In a new study, published today in Nature Communications, a multi-center team led by the Wellcome Sanger Institute, the University of Oslo, Imperial College London and UCL, has mapped for the first time the evolutionary timeline and population distribution of Escherichia coli's protective outer capsule, which is responsible for the bacterium's virulence. The study also shows how targeting the bacterium's protective layer can help treat extraintestinal infections.
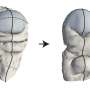
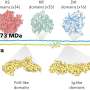
This new work focused on a particular subset of E. coli with a specific capsule—the extracellular barrier that surrounds a bacterium—which scientists have called K1. E. coli with this type of capsule are known to cause invasive diseases such as bloodstream or kidney infections, and meningitis in newborns. This is because this particular cover allows them to mimic molecules already present in human tissues and enter the body unnoticed.
The researchers present evidence that targeting the capsule can be used as the basis of treatment, paving the way to prevent serious E. coli infections.
E. coli is a common cause of urinary tract and bloodstream infections and can cause meningitis in premature and term newborns, with a mortality rate as high as 40%. Furthermore, the rise in hypervirulent and multi-drug resistant E. coli during the last decade means that developing effective strategies to prevent and treat E. coli has now become urgent.
Understanding the bacterium's anatomy and how this plays a role in causing disease is key for the prevention of serious infections. Scientists until now lacked basic knowledge of the prevalence, evolution and functional properties of the K1 capsule, limiting their capacity to combat E. coli infections.
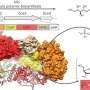
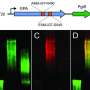
Researchers at the Wellcome Sanger Institute, the University of Oslo, Imperial College London and UCL have now mapped the evolution of this E. coli strain, its prevalence and distribution. Using high-resolution population genomics, whole genome sequencing and advanced computational tools, they analyzed 5,065 clinical samples from different countries and time periods. The data included large collections of samples from the UK and Norway, newly-generated adult and neonatal samples from six countries, such as Brazil, Mexico and Laos among others, and samples from the pre-antibiotic era—from 1932 onwards.
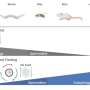
They found that this specifically virulent capsule—K1—actually dates further back in time, approximately 500 years earlier than previously imagined. This highlights the importance of the capsule for the bacterium's survival and the role of the extracellular barrier in the success of E. coli as the main cause of extraintestinal infections.
Dr. Sergio Arredondo-Alonso, lead author of the study from the University of Oslo and the Wellcome Sanger Institute, said, "It was exciting to discover the possibility of reconstructing the evolutionary history of the K1 capsule over the last half millennium, and to see how the capsule genes have been acquired over and over again by many different lineages of this pathogen species over the centuries. As neither the prevalence nor the history of K1 was known, it felt like we entered truly unchartered territory and significantly advanced understanding of this major pathogen species."
The study also shows that 25% of all current E. coli strains responsible for blood infections contain the genetic information needed to develop the K1 capsule. Obtaining a complete evolutionary history of this strain will now allow researchers to understand how bacteria obtain the genetic material responsible for severe virulence in the first place, and analyze ways to combat them.
By using enzymes from bacteriophages, which are viruses that infect and kill bacteria, researchers were able to remove the bacterium's extracellular barrier and make it vulnerable to the human immune system. The researchers demonstrated in in vitro studies using human serum—a liquid part of the blood that is commonly used in laboratory studies—that targeting this capsule can be a way to broadly treat E. coli infection without the use of antibiotics, consistent with previous experimental infections in animals.
Dr. Alex McCarthy, a senior author of the study from Imperial College London, said, "We specifically demonstrated the advances made possible by combining experimental microbiology with population genomics and evolutionary modeling tools, to open a window into translating the findings into future clinical practice. We show that therapeutic targeting of the K1 capsule makes these pathogens more vulnerable to our immune system, and offers the possibility of preventing serious infections. For example, it could help treat newborn babies with meningitis caused by K1 E. coli, which is a rare but dangerous condition associated with high mortality and serious long-term adverse health effects."
Professor Jukka Corander, a co-senior author of the study from the Wellcome Sanger Institute and the University of Oslo, said, "Our research shows the importance of representative genomic surveys of pathogens over time and space. These studies will enable us to reconstruct the evolutionary history of successful bacterial lineages and pinpoint changes in their genetic make-up that can lead to their ability to spread and cause disease. Such knowledge is ultimately providing the basis for designing future interventions and therapies against these pathogens."
More information: Evolutionary and functional history of the Escherichia coli K1 capsule, Nature Communications (2023). DOI: 10.1038/s41467-023-39052-w
Journal information: Nature Communications
Provided by Wellcome Trust Sanger Institute