This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Being noisy in a crowd: How gene interactions shape the evolution of cell-to-cell variation
Biological cells, whether free-living or part of a multicellular organism, have to perform hundreds of functions to survive, such as perceiving their environment, uptaking and metabolizing nutrients, regenerating decayed parts, reproducing themselves, and many more.
The information on how to perform these functions is carried by genes and practically realized through a process called gene expression, through which gene products are made. Gene products work together in what is often represented by a network of interactions known as a gene network.
However, the process of gene expression is subject to randomness, and the expression of each gene in the network is somewhat unpredictable. How do genes in gene networks evolve to cope with this inherent noise while maintaining the function of the gene network? This question is being addressed by the Molecular System Evolution research group at the Max Planck Institute for Evolutionary Biology.
Cells as stochastic machines
The typical textbook representation of genes organized in well-defined networks gives the illusion of the cell operating as a finely tuned, programmed machine, but it is far from it. Studies in single-cell biology revealed that gene expression is inherently a noisy process. Cells with identical genetic backgrounds may express genes in very distinct manners, leading to a form of cellular individuality.
The cell-to-cell variability of the amount of gene products is termed expression noise, and it was shown that this noise spreads and possibly amplifies from one gene to another within the network.
High-throughput genomics studies at the single-cell level further revealed that genes vary extensively by the amount of noise they display. Some are expressed with high accuracy, while others are much less predictable. This extensive variation of noise levels within the genome suggests that expression noise is shaped by natural selection, but how selection acts on genes within networks is largely unknown.
In silico evolution of gene networks
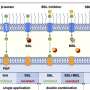
A new study by Nataša Puzović, Tanvi Madaan, and Julien Dutheil from the Max Planck Institute for Evolutionary Biology in Plön investigates the evolution of expression noise in gene regulatory networks with a computational approach. They performed in silico evolution experiments, generating thousands of model gene regulatory network populations and simulated their evolution across multiple generations.
The authors found that a gene's evolved amount of noise highly correlates to its position in the network. Central, highly connected genes regulate other genes and evolve a highly deterministic gene expression. In contrast, peripheral genes at the end of the regulatory chain tend to be more unpredictable.
Although it makes necessary approximations, the model of gene networks allows us to unravel the complex effects of the network structure. The authors show that global network characteristics affect the average noise levels in the network, indicating that the whole network topology should not be disregarded when studying expression noise.
A domino effect
This study demonstrates that selection at the network level to carry out a particular cellular function results in differential selective pressures on individual genes' expression noise and that the network's structure modulates this effect. It proposes noise propagation as the underlying mechanism for the observed variability of expression noise levels in the genomes of organisms. This can be understood as an example of the domino effect: when a central gene is noisy, all other connected genes are affected, and the entire network collapses. On the contrary, a domino falling at the end of the chain has minimal consequences. As a result, the burden of reducing expression noise at the network level is heavier on genes that control other genes.
This study shows that accounting for selection at multiple organizational levels is necessary to understand the evolution of life forms made of many interacting components. It further indicates that natural selection not only acts on the mean expression level, which has been the focus of molecular biology for half a century, but also on its variance and heterogeneity—a dimension that we are only fully starting to unravel with the advent of single-cell omics.
Understanding how genes work
A fundamental goal of biology is discovering how genes work together to create a functioning organism. It is essential to understand how changes in these genes can lead to disease or, conversely, to disease resistance, paving the way to new treatments.
The consideration of individual genes cannot take place in isolation; instead, it is essential to consider and understand their interactions as a system. Likewise, we must not ignore that such systems result from millions of years of evolution.
Computer models, such as those used in the study described here, which integrate our knowledge of both molecular biology and evolutionary processes, are key to this goal.
The paper is published in the journal PLOS Computational Biology.
More information: Nataša Puzović et al, Being noisy in a crowd: Differential selective pressure on gene expression noise in model gene regulatory networks, PLOS Computational Biology (2023). DOI: 10.1371/journal.pcbi.1010982
Journal information: PLoS Computational Biology
Provided by Max Planck Society