This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Researchers uncover the first steps driving antibiotic resistance
Antibiotic resistance is a global health threat. In 2019 alone, an estimated 1.3 million deaths were attributed to antibiotic resistant bacterial infections worldwide. Looking to contribute a solution to this growing problem, researchers at Baylor College of Medicine have been studying the process that drives antibiotic resistance at the molecular level.
They report in the journal Molecular Cell crucial and surprising first steps that promote resistance to ciprofloxacin, or cipro for short, one of the most commonly prescribed antibiotics. The findings point at potential strategies that could prevent bacteria from developing resistance, extending the effectiveness of new and old antibiotics.
"Previous work in our lab has shown that when bacteria are exposed to a stressful environment, such as the presence of cipro, they initiate a series of responses to attempt to survive the toxic effect of the antibiotic," said co-corresponding author Dr. Susan M. Rosenberg, Ben F. Love Chair in Cancer Research and professor of molecular and human genetics, biochemistry and molecular biology and molecular virology and microbiology at Baylor. She also is program leader in Baylor's Dan L Duncan Comprehensive Cancer Center (DLDCCC).
"We discovered that cipro triggers cellular stress responses that promote mutations. This phenomenon, known as stress-induced mutagenesis, generates mutant bacteria, some of which are resistant to cipro. Cipro-resistant mutants keep on growing, sustaining an infection that can no longer be eliminated with cipro."
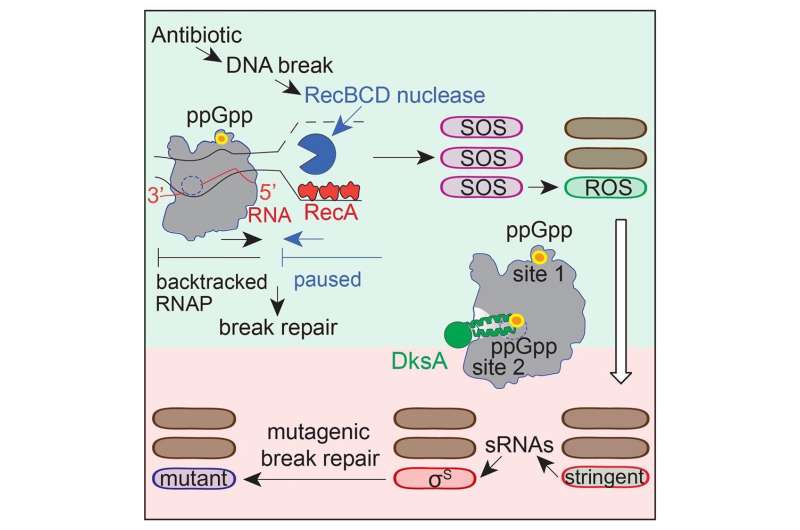
Cipro induces breaks in the DNA molecule, which accumulate inside bacteria and consequently trigger a DNA damage response to repair the breaks. The Rosenberg lab's discoveries of the steps involved in stress-induced mutagenesis revealed that two stress responses are essential: the general stress response and the DNA-damage response.
Some of the downstream steps of the process that leads to increased mutagenesis have been revealed previously by the Rosenberg lab and her colleagues. In this study, the researchers discovered the molecular mechanisms of the first steps between the antibiotic causing DNA breaks and the bacteria turning on the DNA damage response.
"We were surprised to find an unexpected molecule involved in modulating DNA repair," said first author Dr. Yin Zhai, postdoctoral associate in the Rosenberg lab. "Usually, cells regulate their activities by producing specific proteins that mediate the desired function. But in this case, the first step to turn on the DNA repair response was not about activating certain genes that produce certain proteins."
Instead, the first step consisted of disrupting the activity of a protein already present, RNA polymerase. RNA polymerase is key to protein synthesis. This enzyme binds to DNA and transcribes DNA-encoded instructions into an RNA sequence, which is then translated into a protein.
"We discovered that RNA polymerase also plays a major role in regulating DNA repair," Zhai said. "A small molecule called nucleotide ppGpp, which is present in bacteria exposed to a stressful environment, binds to RNA polymerase through two separate sites that are essential for turning on the repair response and the general stress response. Interfering with one of these sites turns off DNA repair specifically at the DNA sequences occupied by RNA polymerase."
"ppGpp binds to DNA-bound RNA polymerase, telling it to stop and backtrack along the DNA to repair it," said co-corresponding author Dr. Christophe Herman, professor of molecular and human genetics, molecular virology and microbiology and member of the DLDCCCC. The Herman lab found the repair-RNA-polymerase connection previously, reported in Nature.
Rosenberg's lab discovered that DNA repair can be an error-prone process. As repair of the broken DNA strands progresses, errors occur that alter the original DNA sequence producing mutations. Some of these mutations will confer bacteria resistance to cipro. "Interestingly, the mutations also induce resistance to two other antibiotic drugs the bacteria have not seen before," Zhai said.
"We are excited about these findings," Rosenberg said. "They open new opportunities to design strategies that would interfere with the development of antibiotic resistance and help turn the tide on this global health threat. Also, cipro breaks bacterial DNA in the same way that the anti-cancer drug etoposide breaks human DNA in tumors. We hope this may additionally lead to new tools to combat cancer chemotherapy resistance."
More information: Yin Zhai et al, ppGpp and RNA-polymerase backtracking guide antibiotic-induced mutable gambler cells, Molecular Cell (2023). DOI: 10.1016/j.molcel.2023.03.003
Journal information: Molecular Cell , Nature
Provided by Baylor College of Medicine