March 3, 2023 report
This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Molecular atlas of spider silk production could help bring unparalleled material to market
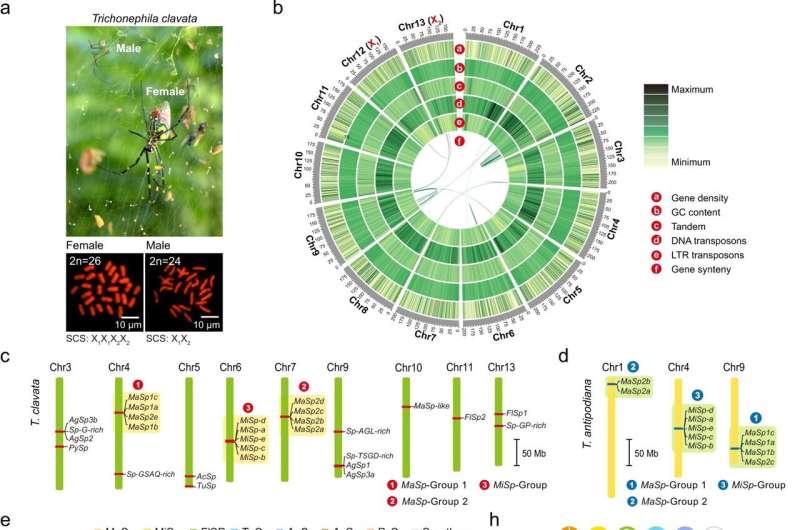
Researchers from Southwest University in China have constructed the entire chromosomal-scale genome assembly and complete spidroin gene set of the golden orb-weaving spider, Trichonephila clavata, known for its especially strong, golden-colored webs.
They attest that their work "Provides multidimensional data that significantly expand the knowledge of spider dragline silk generation…" and the researchers plan on using this new "molecular atlas" to better understand how spiders manufacture their silk.
Published in the journal Nature Communications, the paper details the steps the researchers took, from wild spider capture to multiomic analysis, in revealing the interplay of genes within the spider's major ampullate gland, the gland responsible for producing dragline silk.
Spider dragline silk is a true material marvel with many potential medical and industrial applications. It is lighter and stronger than steel while maintaining an elastic stretching ability that rivals rubber. Unlike many synthetic materials, spider silk is non-toxic, biodegradable, and biocompatible, making it an ideal material for surgical implants, biosensors, and tissue reconstruction.
The only limitation to adopting spider silk as a replacement for a long list of materials we currently use is how difficult it is to manufacture. There was an effort in the past to produce the proteins in goat milk by a company called Nexia, and it worked, but not at a scale required for mass production.
And despite the obvious benefits of spider silk, no one has stepped up to start spider farming on the scale required. Researchers expect that by better understanding silk production at a molecular level in spiders, they will gain practical insights to help bring this unparalleled material to market.
Making a molecular atlas with multiomics
To obtain the genome, the research team used the Oxford Nanopore platform, which can produce the most extended contiguous reads of any gene sequencer, as well as Illumina sequencing machines for more accurate yet shorter read capture lengths and Hi-C for chromosome mapping. By combining these three different sets of genomic data, the researchers were able to bioinformatically reconstruct a detailed model of the spider's chromosomal-scale genome assembly and complete spidroin gene set.
Having this genomic data allows connections to be made between gene expression and, ultimately, the proteins found in spider silk, which is exactly what the researchers did next. The team performed transcriptome (messenger RNA), protein, and metabolite (signaling molecule) analysis of the three segments of the major ampullate gland; the tail, sac, and duct.
Liquid chromatography-mass spectrometry analysis identified 28 proteins: 10 were spidroins, the proteins that make up spider silk, 15 were spider silk-constituting elements, and one was related to venom. With the core components identified, the researchers could rank them in order of intensity-based absolute quantification.
Further analysis allowed them to characterize the specific biological functions of the Tail, Sac, and Duct related to silk production based on the function of genes and gene products. The Tail omics mostly revolved around the synthesis of organic acids, those in the Sac primarily focused on lipid production, and the Duct omics were related to ion exchange and chitin synthesis.
Previous research has found some elements discovered in the current study, but none have put the whole picture together in such a complete and comprehensive way.
More information: Wenbo Hu et al, A molecular atlas reveals the tri-sectional spinning mechanism of spider dragline silk, Nature Communications (2023). DOI: 10.1038/s41467-023-36545-6
Journal information: Nature Communications
© 2023 Science X Network





















