This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Long-standing mystery about mRNAs resolved
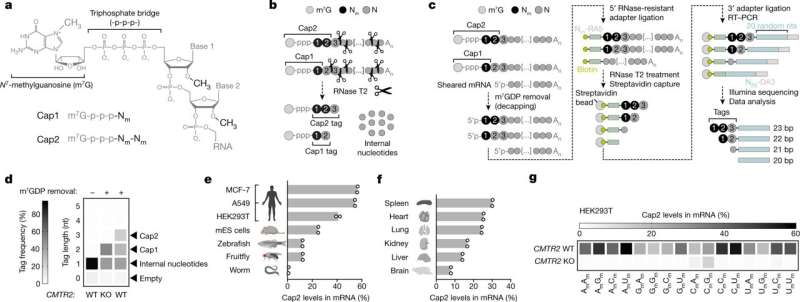
Messenger RNAs (mRNAs) contain chemical marks that are critical for antiviral defense in cells, according to a new study from researchers at Weill Cornell Medicine. The finding solves a 50-year mystery concerning the purpose of these chemical modifications, and suggests that faulty mRNA modification may underlie some autoimmune and inflammatory disorders.
The researchers, whose findings appear Feb. 1 in Nature, discovered that the presence of a common modification, called a methylation, at a particular spot on an mRNA molecule, provides extra protection for the mRNA from antiviral immune mechanisms that might otherwise destroy it.
"We've known since the 1970s that methyl modifications are somehow fundamental to how mRNAs normally work," said study senior author Dr. Samie Jaffrey, the Greenberg-Starr Professor in the department of pharmacology at Weill Cornell Medicine. "So it's very gratifying to finally have this insight into its precise role."
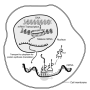
Messenger RNAs are copied from active genes, and are called messengers because they carry protein-making instructions from the DNA in the cell nucleus outward to the main compartment of cells—the cytosol—where they are translated into proteins.
The Jaffrey lab researches the mechanisms that cells use to regulate messenger RNAs—for example, to boost or inhibit their translation into proteins. One type of regulation involves the incorporation of chemical modifications in mRNA. These chemical modifications typically involve methyl modifications.
In previous work, Jaffrey's group developed methods to detect one of these methyl modifications, called methyl adenosine or m6A, which controls mRNA stability in cells. Alterations in m6A can lead to different types of cancer.
However, mRNAs often contain another chemical modification called Cap 2. In their new study, Jaffrey and first author Vladimir Despic, a postdoctoral research associate in the lab, examined this modification whose function has been an enduring mystery.
An mRNA, like the DNA from which it is copied, is a chain of small molecular building blocks called nucleotides. When an mRNA is made, it is "capped" at its first nucleotide with a small organic molecule. The first nucleotide is also modified by the attachment of a small cluster of atoms called a methyl group.
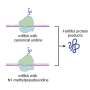
When this type of methylation is present on the first nucleotide, an mRNA is said to have the standard "Cap1" cap, which is known to help shield the mRNA from immune mechanisms that police the cytosol for anything resembling viral RNA.
Intriguingly, some mRNAs acquire an additional methylation at their second nucleotide. Why this additional "Cap2" methylation occurs, and why it is seen on some mRNAs and not others, are questions that have been virtually impossible to answer—mainly because biologists have not had a good method for detecting which mRNAs have Cap2 versus Cap1.
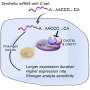
Jaffrey and Despic began their study by developing just such a method, which they call CLAM-Cap-seq. With it, they discovered that the Cap2 methylation can occur on any mRNA, but happens relatively slowly, so that it tends to be found only on mRNAs that have been in the cytosol for longer periods of time.
Ultimately, they found evidence that while Cap1 greatly reduces an mRNA's ability to trigger cellular antiviral mechanisms, Cap2 provides crucial added protection. The researchers observed that when a cell's mRNAs are only of the Cap1 type, these cellular mRNAs activated the cells' inflammatory antiviral mechanisms, even in the absence of viruses.
But too much Cap2 is also bad, the researchers found. When they engineered cells to rapidly incorporate Cap2 into mRNA, they found that the RNAs of invading viruses started to acquire it, shielding them from immune attack and allowing the viruses to grow out of control.
"We think Cap2 methylation occurs slowly, rather than quickly, in order to reduce the chance it will end up cloaking fast-replicating viral RNAs," Jaffrey said.
The findings, apart from resolving the long-standing Cap2 mystery, open up new directions for translational research. One possibility Dr. Jaffrey is now pursuing, he said, is that the dysfunction of the Cap1/Cap2 process underlies some common inflammatory and autoimmune disorders, such as lupus and rheumatoid arthritis, and that correcting this dysfunction could be a new way to treat these disorders.
Another possibility, he said, is to boost antiviral immunity by inhibiting Cap2 in the context of viral infections that otherwise have no good treatment.
"We're also studying the possibility of using Cap2 modifications," Jaffrey said, "to improve mRNA-based therapeutics, including vaccines, by reducing their inflammatory effects in cells."
More information: Vladimir Despic et al, mRNA ageing shapes the Cap2 methylome in mammalian mRNA, Nature (2023). DOI: 10.1038/s41586-022-05668-z
Journal information: Nature
Provided by Cornell University