This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
trusted source
proofread
Have model organisms evolved too far?
A model organism used in laboratories for the past 100 years has evolved so extensively that it may no longer be fit for purpose.
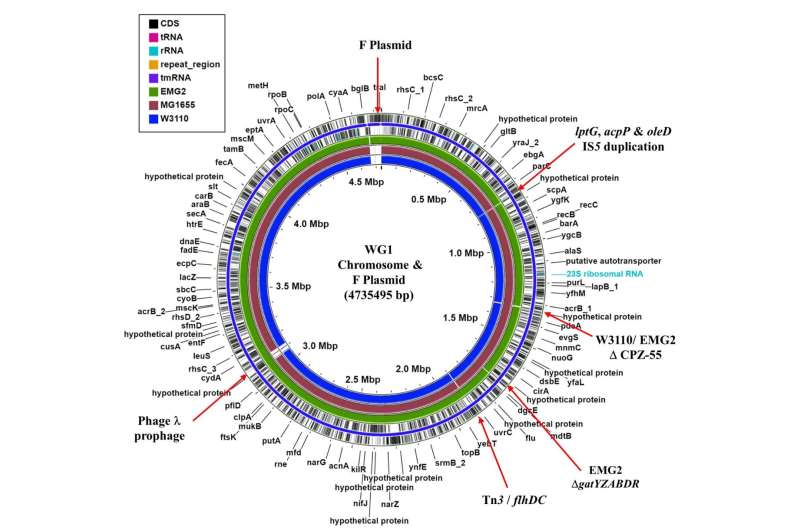
According to a new study, published in Microbial Genomics, the bacterial strain Escherichia coli K-12 has been repeatedly cultured and mutated, resulting in an organism that carries many genetic changes compared to the original isolated bacteria.
The research team, from Aston University, and the Universities of Birmingham and Nottingham, made their discovery after re-examining the early preserved samples and looking at the base sequence of their DNA. They found a large number of differences at the DNA sequence level, and the differences are bigger when they examined currently used stocks that derived from the original samples.
The work underscores the dangers of using one strain as a sole model. It also confirms that bacterial sequences evolve over short time scales, and provides a fascinating insight into the first baby steps of molecular microbiology.
Lead author Doug Browning, of the School of Biosciences at Aston University, said, "The past 10 years have seen a massive amount of bacterial genome sequencing and the picture that is emerging is that bacterial genomes change very fast. This was unimaginable 100 years ago, and, of course, this is why folk back then were quick to adopt the K-12 strain as the model for everything."
The strain of bacteria in the study was originally isolated in 1922 from the feces of a recuperating diphtheria patient at Stanford University, in California. The strain was preserved and over time it, and many derivatives, were distributed to research laboratories around the world for use by researchers looking to understand the workings of living cells at the molecular level.
While the number of genetic variations which have appeared in the intervening decades may sound alarm bells in some research areas, for others it may represent new research opportunities.
Co-author Steve Busby, of the Institute of Microbiology and Infection at the University of Birmingham, said, "Actually the diversity that all this generates can add a new dimension to our understanding. It's often true that things are seldom as they seem, and particularly so if you only study one strain."
More information: Douglas F. Browning et al, Laboratory strains of Escherichia coli K-12: things are seldom what they seem, Microbial Genomics (2023). DOI: 10.1099/mgen.0.000922
Provided by University of Birmingham