This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Decades-old crustaceans coaxed from lake mud give up genetic secrets revealing evolution in action
Human actions are changing the environment at an unprecedented rate. Plant and animal populations must try to keep up with these human-accelerated changes, often by trying to rapidly evolve tolerance to changing conditions.
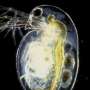
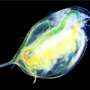
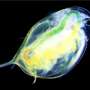
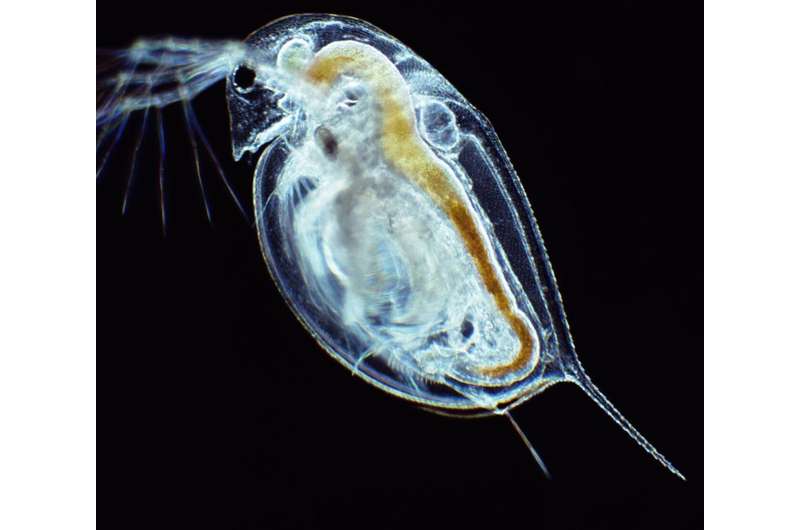
University of Oklahoma researchers Lawrence Weider, professor of biology, and Matthew Wersebe, a biology doctoral candidate, have demonstrated rapid evolution in action by sequencing the genomes of a population of Daphnia pulicaria, an aquatic crustacean, from a polluted lake.
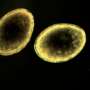
The research, which was conducted as part of Wersebe's doctoral dissertation, was recently published in the Proceedings of the National Academy of Sciences. Wersebe and Weider revived decades-old Daphnia resting eggs from lake sediments, a method known as resurrection ecology, which has been refined in Weider's lab over the past several decades. They then sequenced the entire genomes of 54 different Daphnia individuals from different points-in-time, allowing them to study the genetics and evolution of the population.
The Daphnia were collected from Tanners Lake, located in Oakdale, Minnesota. Tanners Lake has suffered significant salt pollution, stemming from the widespread use of road deicing salts in its watershed.
Daphnia, also known as water fleas, play critical roles in environmental monitoring. For example, they have served as important test organisms in laboratories around the world for over a century because of their sensitivity to many environmental stressors such as chemicals. In nature, Daphnia act as a keystone species in freshwater food webs globally, where they feed on algae to help keep lake and reservoir water clean and serve as a food item for recreational and commercially important fish species.
Wersebe's and Weider's results indicate that rapid adaptation to salt pollution may allow lake Daphnia to persist in the face of anthropogenic salinization, maintaining the food webs and ecosystem services that Daphnia support. However, the ability of these populations to adapt will depend on the speed at which these changes are occurring and the underlying genetic makeup of the impacted populations.
Over the past several years, many researchers have published results defining the scope and scale of lake salinization and recent research has highlighted the ecological impacts. However, to date, the evolutionary implications are not well known. Through their study, Wersebe and Weider reported signatures of natural selection throughout the genome near genes related to osmoregulation and ion regulation, key processes for dealing with high salt. Characterizing clones for salinity tolerance revealed evidence that genetic changes may underlie rapid evolution.
"Work like this is the first step in designing future studies incorporating recent technological advances, such as CRISPR gene editing, allowing the creation of comprehensive genotype-to-phenotype maps and predicting the role that genetic variation plays in creating diverse forms and functions," Wersebe said. "In fact, we found a promising gene that appears not to work properly in the older Daphnia, but a functional copy of the gene is increasing in frequency—true evolution in action."
Future research using these advanced technologies for cutting and pasting the non-functional gene into Daphnia would be one way to better probe the effects that mutations have on complex phenotypic traits like salinity tolerance.
More information: Matthew J. Wersebe et al, Resurrection genomics provides molecular and phenotypic evidence of rapid adaptation to salinization in a keystone aquatic species, Proceedings of the National Academy of Sciences (2023). DOI: 10.1073/pnas.2217276120
Journal information: Proceedings of the National Academy of Sciences
Provided by University of Oklahoma