This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
AI helps scientists decipher cellular structures
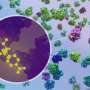
To the untrained eye, a cryo-electron tomogram looks more like traces in sand than the detailed snapshot of a cell it is.
Specialists trained in powerful microscopy techniques like cryo-electron microscopy and tomography can use these images to study the location and shape of cellular organelles and structures of large molecular complexes. As a result, researchers can gain insight into a cell's inner workings, both in healthy and diseased states.
This approach has a major drawback, however. While trained specialists can be very good at recognizing and labeling different cellular structures in tomograms, the process is extremely time-consuming.
This is why the Zaugg, Mahamid, Kreshuk and Diz-Muñoz groups at EMBL Heidelberg have created an artificial intelligence-based method to rapidly and efficiently annotate cellular structures in cryo-electron tomograms. They described this tool in a recent publication in Nature Methods, and have made it openly available for the scientific community to access and use.
DeePiCt (Deep Picker in Context), a deep-learning framework, can recognize and label organelles and molecular complexes substantially faster than the human eye and without human bias, producing richly detailed cellular images (like the one in the circle on the right in the image above).
"DeePiCt—and in particular the trained models that we provide—make it possible for anyone to detect specific particles and structures of interest among the noisy background of their own tomograms. For me, this is one of the best outcomes of our work," said Judith Zaugg. "Without it, you needed to ask a trained specialist for help with annotations, and this took potentially very long. I see DeePiCt as an important step towards enabling high-throughput in cell structural biology."
The DeePiCt framework allows scientists to easily classify cellular structures in tomograms based on where in the cell they are located. This can then be used, for example, to compare the class of ribosomes located on mitochondria with those located on the endoplasmic reticulum. Such analyses have already revealed unknown structural details of how ribosomes bind to these different membranes.
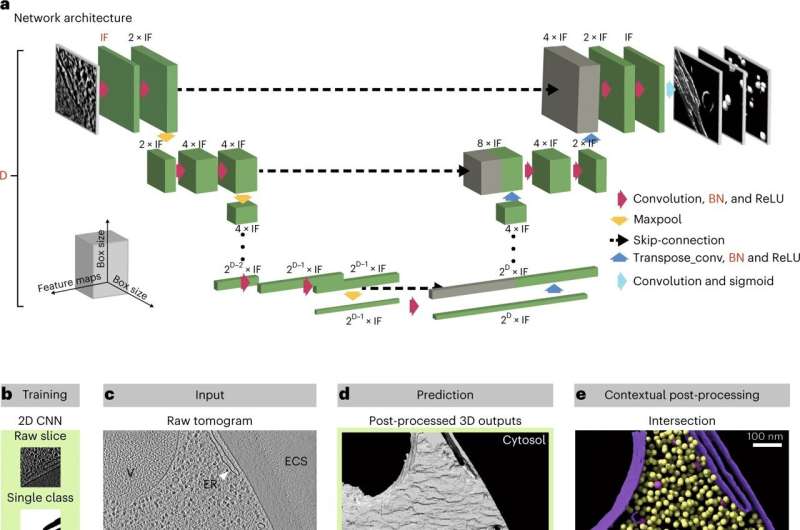
The software combines two types of convolutional neural networks. These are deep learning algorithms that can find patterns and differentiate objects in an image. The first was trained to segment cellular structures like organelles and cytoplasm and operates in 2D slices. The second was trained to segment a particle of interest (e.g. a ribosome) and operates in the three-dimensional space of a tomogram.
Importantly, once the network was trained to recognize a specific particle in a set of tomograms, it could then identify the same particles in new tomograms it had never seen before, including those of cells belonging to a different species. This means DeePiCt can be used by researchers using cryo-electron tomography on many different sample types.
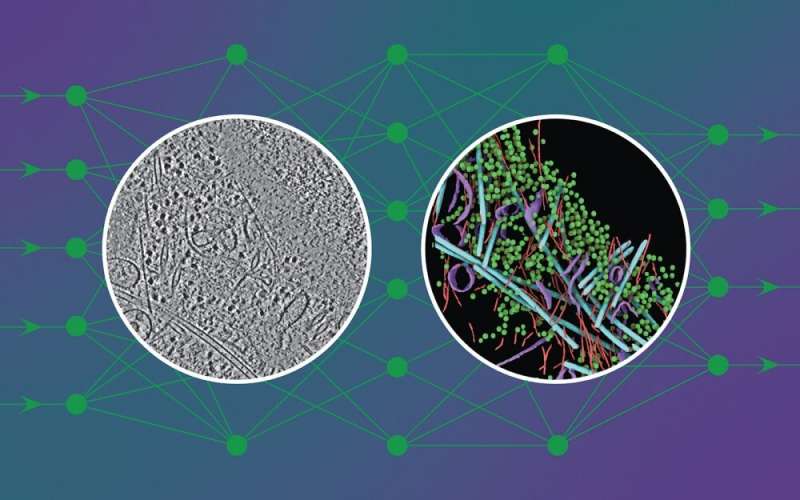

In the image shown, the network was trained to detect four distinct structures (actin, ribosomes, microtubules, and membranes) in cells from three different organisms to predict these structures in an unseen tomogram from a human cell.
"Now we have shown that this works, we are excited about making the software available to the research community," said Julia Mahamid. "We hope that such deep-learning approaches will become established as a gold standard in cryo-electron tomography. And we are providing a set of 20 well-annotated tomograms in the EMBL-EBI archives, which we expect to trigger and support further methods development within the scientific community."
More information: Irene de Teresa-Trueba et al, Convolutional networks for supervised mining of molecular patterns within cellular context, Nature Methods (2023). DOI: 10.1038/s41592-022-01746-2
Journal information: Nature Methods
Provided by European Molecular Biology Laboratory