This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Unraveling the protein map of the cell's mitochondria
Mitochondria are responsible for the energy supply of the organism and fulfill functions in metabolic and signaling processes. Researchers at the University Hospital Bonn (UKB) and the University of Freiburg have gained systematic insight into the organization of proteins in mitochondria.
The protein map of mitochondria represents an important basis for further functional characterization of the powerhouse of cell and thus provide implications for diseases. The study has now been published in Nature.
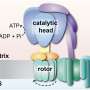
Mitochondria are among the most important cell compartments. They are delimited organelles surrounded by a double membrane. Mitochondria produce the majority of the energy required for all cellular processes. In addition, they take over many other functions in metabolism and provide a signaling surface for inflammatory processes and programmed cell death.
Defects in mitochondria lead to numerous diseases, especially of the nervous system. Therefore, the molecular understanding of mitochondrial processes is of highest relevance for basic medical research. The molecular workers in the cell are usually proteins. Mitochondria can contain around 1,000 or more different proteins.
To execute functions, several of these molecules often work together and form a protein machine, also called a protein complex. Proteins also interact in the execution and regulation of molecular processes. Yet little is known about the organization of mitochondrial proteins in such complexes.
Precision in the analysis of dynamic protein machines
The research groups of Prof. Thomas Becker and Dr. Fabian den Brave at the UKB, together with the research groups of Prof. Bernd Fakler, Dr. Uwe Schulte and Prof. Nikolaus Pfanner at the University of Freiburg, have created a high-resolution image of the organization of proteins in protein complexes, known as MitCOM.
This involved a specific method known as complexome profiling to record the fingerprints of individual proteins at an unprecedented resolution. MitCOM reveals the organization into protein complexes of more than 90 percent of the mitochondrial proteins from baker's yeast. This allows to identify new protein-protein interactions and protein complexes—an important information for further studies.
Quality control in the mitochondrial entry gate TOM as an example
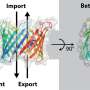
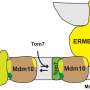
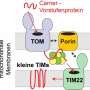
Researchers at UKB in cooperation with Collaborative Research Center 1218, "Regulation of cellular function by mitochondria", have shown how this dataset can be used to elucidate new processes. Mitochondria import 99 percent of their proteins from the liquid portion of the cell, known as cytosol. In this process, a protein machinery called the TOM complex enables the uptake of these proteins through the membrane into the mitochondria.
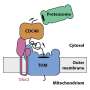
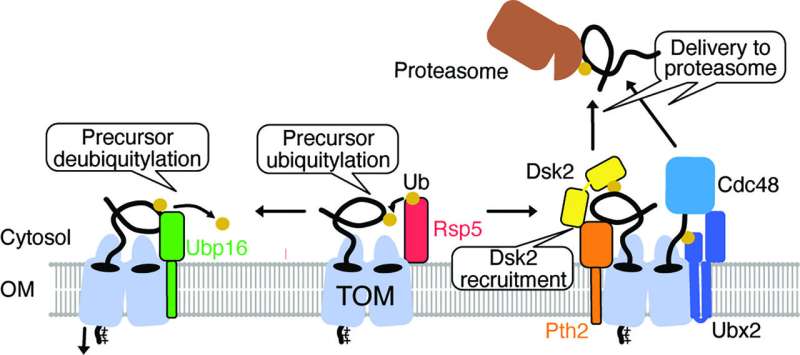
However, it is largely unclear how proteins are removed from the TOM complex when they get stuck during the transport process. To elucidate this, the team led by Prof. Becker and Dr. den Brave used information from the MitCOM dataset. It was shown that non-imported proteins are specifically tagged for cellular degradation.
Research by the Ph.D. student Arushi Gupta further revealed a pathway by which these tagged proteins are subsequently targeted for degradation. Understanding these processes is important because defects in protein import can lead to cellular damage and neurological diseases.
"The example from our study demonstrates the great potential of the MitCOM dataset to elucidate new mechanisms and pathways. Thus, this map of proteins represents an important source of information for further studies that will help us to understand the functions and origin of the cell's powerhouse," says Prof. Becker, director of the Institute of Biochemistry and Molecular Biology at UKB.
More information: Nikolaus Pfanner, Mitochondrial complexome reveals quality control pathways of protein import, Nature (2023). DOI: 10.1038/s41586-022-05641-w. www.nature.com/articles/s41586-022-05641-w
Journal information: Nature
Provided by University of Bonn