This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Stunningly detailed blueprint revealed of viral genome replication machinery

RNA viruses, such as the coronavirus that causes COVID-19, are in a life-and-death race the moment they infect a cell.
These viruses have only minutes to establish their replication machinery inside the host cell before the genetic instructions contained in their vulnerable RNA genomes—which are more fragile than DNA—would otherwise be destroyed by cellular housekeeping. If successful, the virus can go from just a few copies of its RNA genome to a half-million copies incorporated in new infectious particles in less than 12 hours. If not, the virus dies.
In research published online January 24 by the Proceedings of the National Academy of Sciences (PNAS), Morgridge Institute for Research scientists shed new light on these crucial early stages of virus infection and their control. The researchers developed new ways to release viral RNA replication complexes from cells and visualize them in sophisticated ways by cryo-electron microscopy (cryo-EM).
Cryo-EM combines highly advanced imaging with extensive computational analysis to allow scientists to visualize flash-frozen molecules in their native state at molecular to atomic resolution, giving revolutionary insights into biological structure, which can be a powerful foundation for developing therapeutics to thwart disease.
The research team is led led by Paul Ahlquist, director of the Institute's John and Jeanne Rowe Center for Virology and University of Wisconsin-Madison professor of molecular virology and oncology. The team includes scientists Hong Zhan, Nuruddin Unchwaniwala and Johan den Boon, Morgridge investigator and UW-Madison assistant professor of biochemistry Tim Grant, and co-authors Andrea Rebolledo-Viveros, Janice Pennington, Mark Horswill, Roma Broadberry, and Jonathan Myers.
Most microbe and host genes function in large protein complexes that operate as molecular machines. The structures of these critical assemblies, however, have largely been unknown, greatly limiting understanding and control of the relevant processes. In 2017, using an advanced model virus, the Ahlquist lab provided the first full imaging of a viral RNA replication complex and its striking organization.
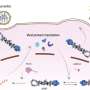
They found the parental viral genomic RNA "chromosome" tightly coiled inside a protective membrane vesicle, whose necked channel to the cytoplasm they discovered to be the site of the actual viral RNA replication machinery—the dynamic, multifunctional engine of genome copying—in a previously unknown, 12-fold symmetric, ringed complex that they named the "crown."
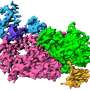
Now, in its new paper in PNAS, the team presents a further leap by revealing the intricate structure of this molecular crown and its component enzyme domains at atomic to near-atomic resolution. These dramatically higher resolution results show how the many distinct functional modules of this replicative engine are arranged, providing an essential basis for working out its assembly, its dynamic operation, and ways to interfere with both.
For comparison, first author Hong Zhan says, "The first visualizations of the crown machinery by our lab in 2017 were like identifying the existence and general outline of a building. The new 2023 resolution is like showing fine details, such as the electrical wiring and door locks."
"In virology," Ahlquist says, "the complexes people have focused on to date mainly were the infectious particles that move between cells, which are relatively easy to purify and study because they release themselves from cells."
"However, most viral replication processes occur in the complex environment within cells," he adds. "This is a new chapter where we've been able to reach inside cells to capture and image in great detail even more intricate viral machinery that carries out the central events of viral replication."
Team member Johan den Boon notes that among other results, they find that "the crown is made of two stacked 12-mer rings of an enormous viral RNA replication protein, whose multiple domains provide all functions required to synthesize new copies of the viral genomic RNA. However, the proteins in the upper and lower rings are in dramatically different conformations, with their constituent domains in different positions relative to each other."
One implication is that the same protein domains operate in distinct ways in the upper and lower rings. Multiple other features underscore that the crown is not a static structure but a sophisticated, active machine that progresses and cycles through a series of movements to carry out its successive activities. Based on this structure and further targeted experiments, the Morgridge team is elucidating the crown's functions and conformational gymnastics.
Another valuable finding from these studies is that the lower 12-mer ring is an assembly precursor that forms prior to the actual steps of RNA replication. This "proto-crown" precursor then recruits the viral genomic RNA template and other components to initiate synthesis of new RNAs, and serves as a base to assemble the mature, double ring replication complex.
Growing evidence suggests that the crown not only synthesizes new copies of the viral RNA genome, but also helps deliver these new genomes into downstream processes of gene expression and assembly of new infectious viral particles. The crown thus appears to provide major functions for organizing many critical phases throughout infection.
"Just slowing down the assembly and function of RNA replication complexes is enough to kill these viruses," Ahlquist says. "These new results provide a strong basis for finding new ways to do that."
Ahlquist and other team members praise the UW-Madison Cryo-EM Research Center (CEMRC) and its leadership as crucial to their progress. CEMRC is making this valuable technology accessible to scores of scientists across the UW-Madison campus, and as a national center, far beyond. Led by biochemistry Professor Elizabeth Wright, CEMRC provides advanced capabilities in essentially all forms of cryo-EM imaging.
Emerging results from the Morgridge group and other researchers indicate that the principles revealed by these studies are evolutionarily ancient, and that similar crown-like complexes are central to the replication of most, if not all, RNA viruses in this large class. This includes the COVID-19 SARS-CoV-2 coronavirus and many other pathogens.
Accordingly, the conserved underlying principles might serve as the basis for developing more powerful, broad-spectrum antiviral strategies that could inhibit infection by not just one but whole groups of viruses, Ahlquist says.
More information: Hong Zhan et al, Nodavirus RNA replication crown architecture reveals proto-crown precursor and viral protein A conformational switching, Proceedings of the National Academy of Sciences (2023). DOI: 10.1073/pnas.2217412120
Journal information: Proceedings of the National Academy of Sciences
Provided by Morgridge Institute for Research