This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
'Friend or foe' bacteria kill their algal hosts when coexisting is no longer beneficial
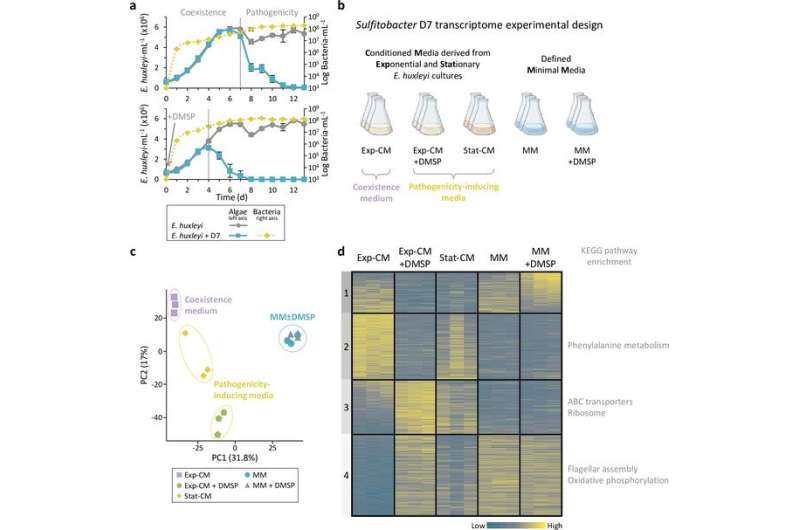
Scientists have detailed a lifestyle switch that occurs in marine bacteria, in which they change from coexisting with algae hosts in a mutually beneficial interaction to suddenly killing them. The results are published today in eLife.
Details of this lifestyle switch could provide new insights into the regulation of algal bloom dynamics and its impact on large-scale biogeochemical processes in marine environments.
Single-celled algae, known as phytoplankton, form oceanic blooms that are responsible for around half of the photosynthesis that occurs on Earth, and form the basis of marine food webs. Therefore, understanding the factors controlling phytoplankton growth and death is crucial to maintaining a healthy marine ecosystem. Marine bacteria from the Roseobacter group are known to pair up and coexist with phytoplankton in a mutually beneficial interaction. The phytoplankton provide the Roseobacter with organic matter useful for bacterial growth, such as sugar and amino acids, and the Roseobacter in return provides B-vitamins and growth-promoting factors.
However, recent studies have revealed that Roseobacters undergo a lifestyle switch from coexistence to pathogenicity, a state in which they kill their phytoplankton hosts. A chemical compound called DMSP is produced by the algae and is hypothesized to play a role in this switch.
"We have previously identified that the Roseobacter Sulfitobacter D7 displays a lifestyle switch when interacting with the phytoplankter Emiliania huxleyi," states first author Noa Barak-Gavish, a Ph.D. graduate in the Department of Plant and Environmental Sciences, Weizmann Institute of Science, Israel. "However, our knowledge about the factors that determine this switch was still limited."
To characterize this lifestyle switch, Barak-Gavish and colleagues performed a transcriptomics experiment, allowing them to compare the genes that are differentially expressed by Sulfitobacter D7 in coexistence or pathogenicity stages.
Their experimental setup demonstrated that Sulfitobacter D7 grown in a pathogenicity-inducing medium have a higher expression of transporters for metabolites such as amino acids and carbohydrates than those grown in a coexistence medium. These transporters serve to maximize the uptake of metabolites released from dying Emiliania huxleyi (E. huxleyi) . Furthermore, in pathogenic Sulfitobacter D7, the team observed an increased activation of flagellar genes that are responsible for the movement of the bacteria. These two factors allow Sulfitobacter D7 to utilize an "eat-and-run" strategy, where they beat competitors to the material released upon E. huxleyi cell death and swim away in search of another suitable host.
The team confirmed the role of DMSP in bringing about the switch to this killer behavior by mapping the genes activated in Sulfitobacter D7 in response to the presence of DMSP and other algae-derived compounds. However, when only DMSP was present, the lifestyle switch did not occur. This implies that although DMSP mediates the lifestyle switch, it is also dependent on the presence of other E. huxleyi-derived infochemicals—compounds that are produced and used by organisms to communicate.
DMSP is an infochemical produced by many phytoplankton, so it is likely that the other required infochemicals allow the bacteria to recognize a specific phytoplankton host. In natural environments, where many different microbial species exist together, this specificity would ensure that bacteria only invests in altering gene expression and its metabolism when the correct algal partner is present.
The study also uncovers the role of algae-derived benzoate in Sulfitobacter D7 and E. huxleyi interactions. Even in high concentrations of DMSP, benzoate functions to maintain the coexistence lifestyle. Benzoate is an efficient growth factor and is provided by E. huxleyi to Sulfitobacter D7 during coexistence. The authors propose that as long as Sulfitobacter D7 benefits from coexistence by receiving materials for growth, it will maintain the mutualistic interaction. When less benzoate and other growth substrates are provided, the bacteria undergoes the lifestyle switch and kills its phytoplankton host, swallowing up any remaining useful materials.
The exact mechanism of Sulfitobacter D7 pathogenicity against E. huxleyi remains to be discovered, and the authors call for further work in this area. The cellular machinery Type 2 secretion system—a complex that many bacteria use to move materials across their cell membrane—is more prevalent in Sulfitobacter D7 compared to other Roseobacters, hinting at a unique method of pathogenicity that requires further investigation.
"Our work provides a contextual framework for the switch from coexistence to pathogenicity in Roseobacter-phytoplankton interactions," concludes senior author Assaf Vardi, a professor in the Department of Plant and Environmental Sciences, Weizmann Institute of Science. "These interactions are an underappreciated component in the regulation of algal bloom dynamics and further study in this area could provide insights into their impact on the fate of carbon and sulfur in the marine environment."
More information: Noa Barak-Gavish et al, Bacterial lifestyle switch in response to algal metabolites, eLife (2023). DOI: 10.7554/eLife.84400
Journal information: eLife
Provided by eLife