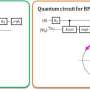
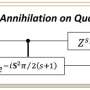
Quantum algorithm of the direct calculation of energy derivatives developed for molecular geometry optimization
In recent years, research and development on quantum computers has made considerable progress. Quantum chemical calculations for electronic structures of atoms and molecules are attracting great attention as one of the most promising applications for quantum computers. In order to utilize quantum chemical calculations for chemistry and related fields, it is essential to develop geometry optimization methods to find the most stable structure for molecules. Geometry optimization requires calculations of energy derivatives with respect to nuclear coordinates of molecules.
The finite difference method is one approach for energy derivative calculations. On a classical computer, calculations based on this method for one-dimensional systems require at least two evaluations of the energy. Previous research has shown that a quantum computer, in contrast, requires only a single query to calculate the energy derivatives based on the finite difference method, regardless of the number of degrees of freedom. However, quantum circuits relevant to quantum algorithms capable of performing energy derivative calculations have not been implemented.
A research group including Dr. Kenji Sugisaki, Professor Kazunobu Sato, and Professor Emeritus Takeji Takui from the Graduate School of Science at Osaka Metropolitan University has successfully extended the quantum phase difference estimation algorithm, a general quantum algorithm for the direct calculations of energy gaps, to enable the direct calculation of energy differences between two different molecular geometries. This allows for the computation, based on the finite difference method, of energy derivatives with respect to nuclear coordinates in a single calculation.
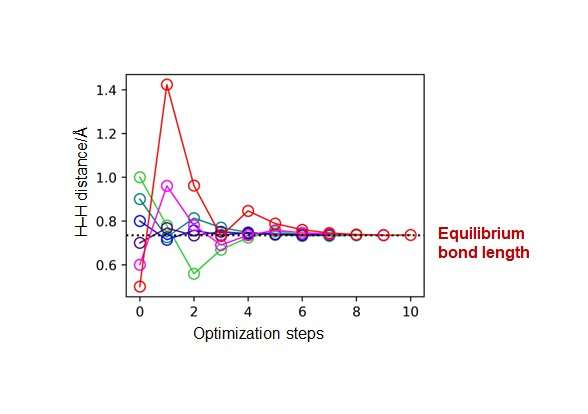
Furthermore, the research group has applied the developed energy derivative calculations to execute geometry optimizations of H2, LiH, BeH2, and N2 molecules without calculating the total energies, demonstrating the usefulness of the developed method. The group also discussed how quantum circuits can be assembled according to different degrees of freedom of the molecules.
This research is the latest in a series of the researchers' articles on quantum chemical calculations on quantum computers. "Our latest findings bring us one step closer to applying quantum chemical calculations on a quantum computer to real-world problems," said Dr. Sugisaki.
"Since energy derivative calculations are used for not only molecular geometry optimizations but also various calculations for molecular properties, the application of our method is expected to play a very important role in a wide range of related fields, such as in silico drug discovery/design and materials development."
The study is published in The Journal of Physical Chemistry Letters.
More information: Quantum Algorithm for Numerical Energy Gradient Calculations at the Full Configuration Interaction Level of Theory, The Journal of Physical Chemistry Letters (2022). DOI: 10.1021/acs.jpclett.2c02737
Journal information: Journal of Physical Chemistry Letters
Provided by Osaka Metropolitan University