Examining previously unknown molecular mechanisms for environmental adaptation in plants
The Plant Physiology and Biochemistry research team at the University of Konstanz has discovered previously unknown molecular mechanisms by which plants adapt to their environment—important basic knowledge in times of climate change
Plants are exposed to constant environmental changes, and their survival depends on their ability to sense and adapt to environmental stimuli. Protein molecules in the cell membrane play a crucial role in coordinating extracellular signals and intracellular reactions. The Plant Physiology and Biochemistry research team (University of Konstanz) now succeeded in identifying two deubiquitinating enzymes involved in the molecular mechanism of this adaptation process. The study has been published in the current issue of Nature Communications.
The amount of protein molecules is crucial
In its adaptation process, the cell needs to sense, for example, nutrients or pathogens in its environment. That is the job of special protein molecules—the transporters and receptors, located on the cell membrane separating the inside of the cell from the outside world. They are produced as well as degraded in the cell. Decisive for the plant's signal perception is the amount of these molecules.
The small signal protein ubiquitin hooks on other proteins and thus ensures that they are degraded. At the same time, there are deubiquitylating enzymes that can reverse this effect by removing ubiquitins.
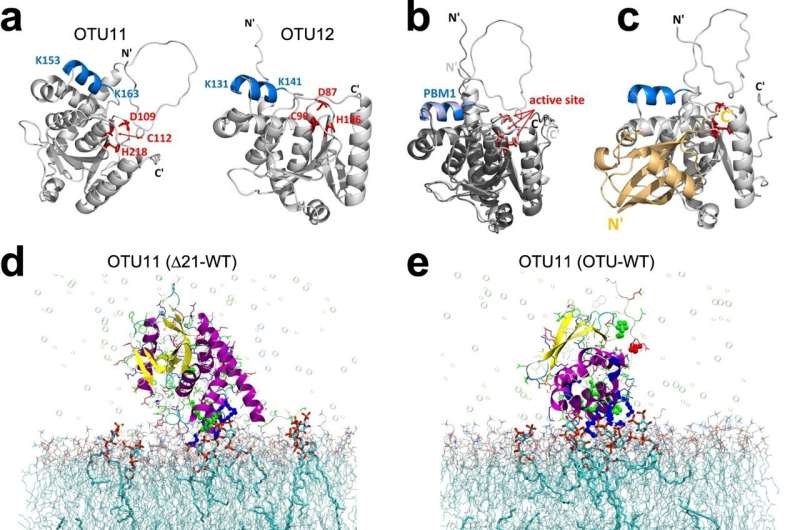
Biology professor Erika Isono's team analyzed 18 of these deubiquitylating enzymes in the model plant Arabidopsis thaliana. In cooperation with Karin Hauser, Michael Kovermann and Christine Peter from the Department of Chemistry, the team found two of these enzymes, designated OTU11 and OTU12, which are localized at the cell membrane and are actually involved in regulating the amount of cell membrane proteins.
Lead author Karin Vogel explains the biochemical mechanism: "OTU11 and OTU12 can shorten certain ubiquitin chains. This influences the degradation of the modified proteins." The biologist also discovered how they are activated: by binding to negatively charged lipids on the cell membrane.
Previously unknown form of activity adjustment
This means that the activity of these enzymes is tightly controlled. Several activation mechanisms for deubiquitylating enzymes are already known. The mechanism reported here is a previously unknown form of activity adaptation. This regulation is so crucial because the effect of deubiquitylating enzymes can have major consequences for the function of the cell.
"Our discovery shows that the deubiquitylating enzymes do not become active until they arrive at the membrane, where the lipids are located. This fits perfectly with the intracellular localization and function of the two enzymes," says Erika Isono.
Model plants are used in basic biological research to investigate fundamental biochemical and molecular biological mechanisms. The long-term goal is to optimize agricultural yields, which is particularly important in times of climate change, as plant growing conditions can change as a consequence. Erika Isono says, "It's important that we understand at the molecular level how plants respond to the environment. The ubiquitin-dependent signaling pathway probably plays an important role in this process."
More information: Karin Vogel et al, Lipid-mediated activation of plasma membrane-localized deubiquitylating enzymes modulate endosomal trafficking, Nature Communications (2022). DOI: 10.1038/s41467-022-34637-3
Journal information: Nature Communications
Provided by University of Konstanz