Using mathematics to treat malaria

The global effect of malaria is devastating. In 2020, there were more than 240 million cases and over 600,000 deaths, mostly in lower income countries in Africa. The most vulnerable are children under the age of five.
The World Health Organization has set ambitious targets for eliminating malaria by 2030. However, these malaria elimination targets hinge on having effective treatments—a difficult feat given the parasite that causes malaria, Plasmodium falciparum, is developing resistance to the different antimalarial drugs in use.
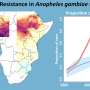
The parasite, which is spread by the Anopheles mosquito, is capable of developing some resistance to all currently available antimalarials, including the most pivotal artemisinin derivatives that are the most effective, and this resistance is spreading.
In Africa, where by far the majority of malaria cases and deaths occur, the antimalarial drug sulfadoxine-pyrimethamine (SP) is used as a preventive treatment for infants, children and during pregnancy. However, we know resistance to SP is spreading as we can track a marker of resistance in the parasite itself—a genetic mutation in the dihydropteroate synthase (pfdhps) gene.
This mutation can undermine the effectiveness of SP and, without the protection of SP during pregnancy, babies are at increased risk of premature birth and low birth weight.
There is an urgent need to be able to track and predict the levels of SP resistance so as to guide its use and indicate to health agencies whether an alternative antimalarial should be used in particular regions. Indeed, as resistance to other antimalarials spreads, so too will agencies want to know how resistance to other antimalarials is varying across different regions and at different times.
So, how can we do this? With mathematics.
Working with the WorldWide Antimalarial Resistance Network (WWARN), based at the University of Oxford and funded by the Bill and Melinda Gates Foundation, our team of international researchers have developed sophisticated mathematical models to reliably predict genetic resistance to SP in Africa in place and in time.
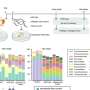
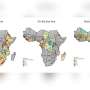
In our research, newly published in PLOS Computational Biology, we used data on the prevalence of the pfdhps mutations in the open-access WWARN repository to develop a statistical model. This allowed us to map the estimated prevalence of the SP resistance marker in Africa from 1990 to 2020.
These maps fill in the gaps where no information is available and can be used by health agencies to guide new polices about where and when the drug is appropriate to use—providing much needed insight about where SP can be used as part of preventive treatments.
SP for malaria prevention in infants is recommended only in areas where the prevalence of a particular pfdhps mutation (540E) is below 50%. For SP use during pregnancy, it has been suggested the SP doesn't have a protective effect in areas where the prevalence of another pfdhps mutation (581G) exceeds 10%.
However, pfdhps mutation prevalence data in Africa are extremely heterogenous and scattered, with data completely missing from many areas. This is where the outputs of our mathematical model can help.
These maps will have a direct public health impact. Spatial information like this on the spread of antimalarial resistance is critical for health organizations to prioritize surveillance measures, and plan control and elimination efforts.
According to our predictive maps, the high prevalence of the pfdhps540E mutation was restricted to East and Southeast Africa, which is reassuring for the continued use of SP for infants in West Africa. However, continuous monitoring will be needed because we predict that the pfdhps540E mutation is spreading.
We have also identified several areas where the prevalence of pfdhps581G exceeded 10%. More data on the distribution of pfdhps581G in these areas will need to be collected to guide SP use during pregnancy.
This research underlines the importance of data sharing and the effectiveness of employing predictive modeling to highlight areas of concern for SP resistance that extend beyond national borders.
The WWARN SP Molecular Surveyor database has been built to provide users with a standardized, current source of information on resistance marker distribution, a model that could be expanded to all validated markers associated with antimalarial resistance.
Given the emergence of resistance to the artemisinin antimalarials in Rwanda, Uganda, Eritrea and Ghana, our approach could be easily extended to better understand the evolution of this new and significant threat.
More information: Jennifer A. Flegg et al, Spatiotemporal spread of Plasmodium falciparum mutations for resistance to sulfadoxine-pyrimethamine across Africa, 1990–2020, PLOS Computational Biology (2022). DOI: 10.1371/journal.pcbi.1010317
Journal information: PLoS Computational Biology
Provided by University of Melbourne