Chemical-free re-staining of tissue using deep learning
Tissue-based diagnosis of diseases relies on the visual inspection of biopsied tissue specimens by pathologists using an optical microscope. Before putting the tissue sample under a microscope for inspection, special chemical dyes are applied to the sample for staining, which enhances the image contrast and brings color to various tissue constituents. This chemical staining process is laborious and time-consuming, performed by human experts. In many clinical cases, in addition to the commonly used hematoxylin and eosin (H&E) stain, pathologists need additional special stains and chemicals to improve the accuracy of their diagnosis. However, using additional tissue stains and chemicals is slow and results in extra costs and delays.
In a recent work published in ACS Photonics, UCLA researchers developed a computational approach powered by artificial intelligence to virtually transfer (re-stain) images of tissue already stained with H&E into different stain types without using any chemicals. In addition to significantly saving expert technician time, chemical staining-related costs and toxic waste generated by histology labs, this virtual tissue re-staining method is also more repeatable than the staining performed by human technicians. Furthermore, it saves the biopsied tissue for more advanced diagnostic tests to be performed, eliminating the need for a second unnecessary biopsy.
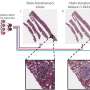
Previous methods to perform virtual stain transfer faced one major problem: a tissue slide can be stained once with one type of stain, and washing away the existing stain and putting a new chemical stain is very difficult and rarely practiced in clinical settings. This makes acquiring paired images of different stain types very challenging, which is an essential part of deep learning-based image translation methods.
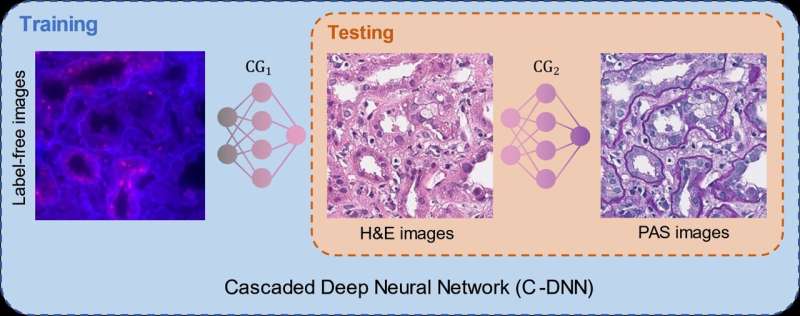
To alleviate this problem, the UCLA team demonstrated a new virtual stain transfer framework using a cascade of two different deep neural networks working together. During the training process, the first neural network learned to virtually stain autofluorescence images of unstained tissue into H&E stain, and the second neural network that is cascaded to the first one learned to perform stain transfer from H&E into another special stain (PAS). This cascaded training strategy allowed the neural networks to directly exploit histochemically stained image data on both H&E and PAS stains, which helped perform highly accurate stain-to-stain transformations and virtual re-staining of existing tissue slides.
This virtual tissue re-staining method can be applied to various other special stains used in histology and will open up new opportunities in digital pathology and tissue-based diagnostics.
This research was led by Dr. Aydogan Ozcan, Chancellor's Professor and Volgenau Chair for Engineering Innovation at UCLA Electrical & Computer Engineering and Bioengineering. The other authors of this work include Xilin Yang, Bijie Bai, Yijie Zhang, Yuzhu Li, Kevin de Haan, and Tairan Liu. Dr. Ozcan also has a faculty appointment in the surgery department at UCLA David Geffen School of Medicine and is an associate director of the California NanoSystems Institute (CNSI).
More information: Xilin Yang et al, Virtual Stain Transfer in Histology via Cascaded Deep Neural Networks, ACS Photonics (2022). DOI: 10.1021/acsphotonics.2c00932
Journal information: ACS Photonics