Rapid, single-cell analysis of microbiotas now possible
A single-cell method developed by RIKEN biophysicists, that can rapidly classify hundreds of thousands of bacteria according to species, promises to be an invaluable tool for discovering how gut, skin, ocean and soil microbes vary with changing conditions.
There has been an explosion in awareness of the various critical roles that bacteria in our guts play in health and disease. Biologists want to explore how the make-up of the gut microbiota affects their hosts, but methods for classifying bacteria according to species provide only very rough results.
Now, Katsuyuki Shiroguchi of the RIKEN Center for Biosystems Dynamics Research and his co-workers have devised a truly single-cell method that can accurately and rapidly characterize communities consisting of hundreds of thousands of bacteria. Their results appear in Nature Communications.
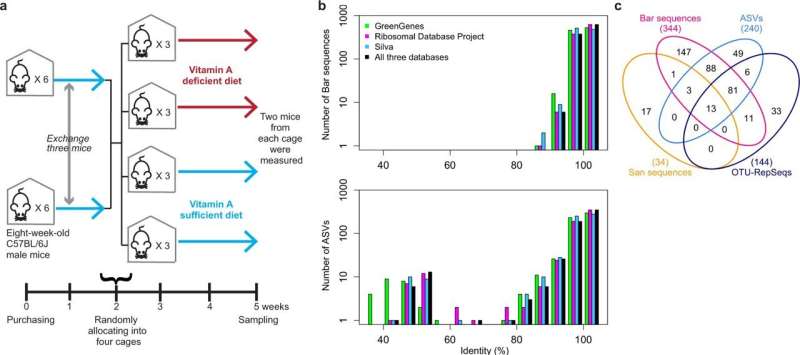
Their method involves encasing individual bacteria in water-in-oil droplets and then labeling a stretch of bacterial DNA coding 16S ribosomal RNA (rRNA) with a unique DNA barcode. This barcode is "read" during sequencing, allowing individual bacteria to be identified. And since each species of bacteria has basically its own 16S rRNA, the method can determine the number of bacteria in a particular species.
The team demonstrated the power of their technique by using it to see how diets deficient in vitamin A affected the gut microbiota of mice. They counted the bacteria at a distal location in the cecum—the entrance to the large intestine—of mice that had been fed a vitamin-A-deficient diet and control mice that had been fed a normal diet. The results revealed that only one species of gut bacteria was significantly affected. "We counted more than 200 different bacterial species in the cecum, but the amount of only one species changed significantly," says Shiroguchi. "This tells us that our method can detect changes in just one species of bacteria."
He is excited about the potential of the method to assess the effects of recently developed bacterial therapies on the gut microbiome. "If we knew that this bacterial species is useful for addressing a specific health problem, doctors could prescribe it, but it hasn't been possible to accurately measure the change in the number of cells in the gut microbiome until now," explains Shiroguchi. "We want to contribute to such medical applications by accurately measuring the number of cells before and after a bacterial therapy."
The team now plans to apply their method to other communities of bacteria besides those in the gut. "We can use it to measure the microbiotas from the soil, sea, atmosphere and skin," he notes.
More information: Jianshi Jin et al, High-throughput identification and quantification of single bacterial cells in the microbiota, Nature Communications (2022). DOI: 10.1038/s41467-022-28426-1
Journal information: Nature Communications
Provided by RIKEN