Online platform designed to improve reproducibility, scientific collaborations

For centuries, scientists relied on a pen or pencil and trusty lab notebook to make sure their experiments could be understood and replicated by colleagues. Now, as experiments may involve dozens of steps and hundreds of materials, produce gigabytes of data that require supercomputers to process and are shared with collaborators around the globe, the lab notebook may no longer suffice.
In a recent study, the researchers report on the development of an online platform that can help genomic researchers track experiments from conception to publication, keeping exacting records for quality control purposes and easing potential reproducibility efforts.
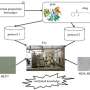
The system, named Platform for Epi-Genomic Research, or PEGR, is designed as a tool to help experimental life sciences laboratories—or wet-labs—keep track of highly complex operations and turn raw data into scientific insights. For example, rather than relying on painstaking hand-written notes, PEGR incorporates two dimensional barcodes—called quick response, or QR codes—to electronically collect and track detailed information on samples as they advance along the experimental process.
The efficiency of PEGR can improve reproducibility, a key reason for the development of the tool, according to William Lai, an assistant research professor, Cornell University, and formerly an assistant research professor in biochemistry and molecular biology, Penn State. Reproducibility, a critical step in the scientific process, requires scientists to check their work to make sure it is accurate, safe and functional in real world applications.
"It's already well recognized that reproducibility is an issue not only in the life sciences, but across all STEM (science, technology, engineering and medical) fields," said Lai. "There has been story after story of research teams that claimed to discover something and then, a few years later, we find that no one has reproduced those results outside of the lab that generated the findings. PEGR is an approach to getting a handle on tracking experimental processes—what a user is using and when are they using it—so that we can improve reproducibility."
Because it is an online platform, PEGR can connect scientists across the world to facilitate reproducibility efforts. The platform also addresses the rapid development of equipment for genomic research—including robotic sampling and high throughput sequencers that can run many experiments simultaneously—that create vast amounts of data, according to Danying Shao, research and development engineer for the Institute for Computational and Data Science's (ICDS) Research Innovations with Scientists and Engineers, or RISE, team.
"There is no doubt a data explosion is happening in bioinformatics," said Shao, who helped design the platform. "Big data sets are being generated at an unprecedented pace. For example, a single sample can generate gigabytes of data. And, when we are sequencing hundreds of samples, you can see that we can get to the level where we are creating terabytes of data."
According to the researchers, who published details about the system in Genome Biology, PEGR is integrated with the Galaxy platform, an open-source scientific workflow system. PEGR is designed to track the sample and sequencing experiment, manage the processing of the data and then produce reports and visualizations of the experimental results.
In the first few runs with the platform, the researchers are already experiencing early benefits.
"Just as an example, recently a technician was experiencing a string of failed experiments, so we went into PEGR and by examining the experimental metadata, we realized that they were using a bad batch of a certain chemical," said Lai. "Now, historically, the process to find the cause of the failing experiments could have dragged on for months—if not, a year or two—instead of finding the source right away."
RISE-powered research
According to Chuck Pavloski, RISE team lead, the PEGR project is just one example of how members of RISE are helping Penn State researchers, as well as the research community at large. Pavloski likens the team a bond between researchers with computational tools and expertise that can expand the power of science to tackle important scientific and societal challenges.
"The RISE engineers are effectively the glue between science and today's computational needs," said Pavloski. "In other words, they allow the scientists to be what they're good at. We act much the same way as a staff scientist works at a national lab, paving the way so that our scientists can explore their fields and pursue their research ideas."
This RISE-powered partnership can help scientists with traditional computational research questions, such as providing guidance on best practices for using Penn State's Roar supercomputer to offering ways to optimize and improve code, but the team can also apply their own deep understanding of academic research to collaborate with scientists on cutting-edge, interdisciplinary projects.
"The RISE team is made up of master's and Ph.D.-level scientists who have a deep understanding of how science works, but work outside of their fields all the time," said Pavloski. "For example, we might have a trained meteorologist who also work on biochemistry or genomic projects, or we have engineers on our team who may be help scientists in astronomy or biochemistry."
This interdisciplinarity offers another advantage. RISE team members can leverage best computational science practices in one field to investigations in other fields or disciplines.
"We also provide a strong link to new technologies, such as using artificial intelligence techniques, or exploring the use of graphic processing unit— or GPU—computation in a research project," Pavloski said.
RISE engineers are also working with data visualization specialists to help scientists create compelling visualizations for their work, as well as take advantage of new immersive technologies, such as virtual reality and augmented reality, to explore data in deeply engaging ways.
Future uses
The researchers hope that PEGR, which is open source, could produce benefits across the scientific enterprise, saving time, money and headaches, and lead to everything from richer understanding of the genome to better medical treatments reaching patients faster.
In the future, the researchers may explore whether the online platform can be expanded outside of use in wet-labs and for use in translational science, which would help scientists bring treatments and solutions to the real world.
"This platform was originally designed around basic research, but we're actively working to move it into the translational biomedical field in the future," said Lai.
More information: Danying Shao et al, PEGR: a flexible management platform for reproducible epigenomic and genomic research, Genome Biology (2022). DOI: 10.1186/s13059-022-02671-5
Journal information: Genome Biology
Provided by Pennsylvania State University