Multi-scale imaging confirms protein's role in neuronal structure, dynamics
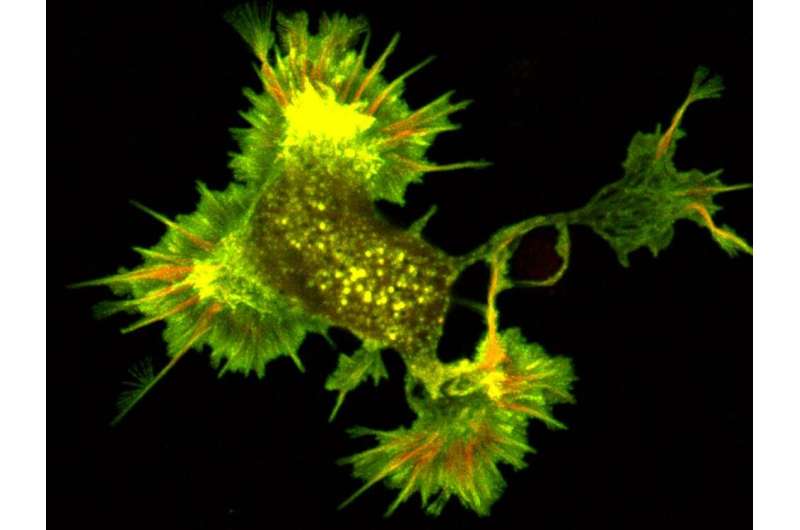
Protein structures are typically determined by studying them in their purified form, outside the busy inner workings of the cell, and because of this, their biological relevance is often called into question. In a new study by Penn State College of Medicine researchers, the long-observed protein structure cofilactin, a form of the filamentous protein actin that contains numerous connections to cofilin proteins, was shown to be a major component of neuronal growth cone filopodia—small dynamic "antennae" at the tips of growing neurons.
"The effects of cofilin on actin structure and its physical properties have been studied for more than 20 years, but now we can confidently see that this structural alteration serves some biological function in the cell," said Matt Swulius, assistant professor of biochemistry and molecular biology. "We're still trying to determine the mechanistic details of its function, but we have strong evidence that cofilactin regulates the flexibility of searching filopodia."
According to Swulius, understanding how filopodial dynamics are controlled at the molecular level could open therapeutic avenues into nerve regeneration as well as some developmental diseases. His lab is studying how the proteins fascin and cofilin function together to regulate the structure and movement of neuronal filopodia as they navigate their environment to eventually form cellular connections.
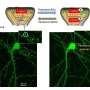
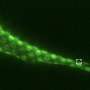
Swulius and team used fluorescence microscopy to reveal the distribution of fascin and cofilin along neuronal filopodia. They found that fascin tended to accumulate at the tip of the protrusion, while cofilin was more prevalent towards the base. Through these fluorescence studies, the team uncovered that a transition region exists toward the middle of the filopodium. In this transition zone, a mixture of fascin and cofilin existed that was distinct from the fascin-rich tip and the cofilin-rich base.
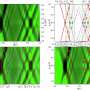
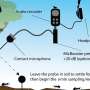
Next, they used cryo-electron tomography, a state-of-the-art three-dimension imaging method, to observe neuronal filopodia in their natural state at molecular resolution. They discovered that actin filaments in the filopodial bundles can switch between interactions with fascin and cofilin, which results in a change in the filaments' helical pitch and packing order within filopodial actin bundles. The transition zones, where both types of filaments were present, were also the areas where the filopodia had curvature, or the ability to move in their environment. They propose that this happens when cofilactin filaments permeate the fascin-linked region and disrupt the fascin cross-links and as a result, increase the flexibility of the filopodial bundle. The results were published in Nature Communications on May 4.
Swulius said there's still a lot to uncover to better understand how cofilin regulates the flexibility of filopodia. They are working to modulate the genetic expression of fascin and cofilin to learn more how both proteins contribute to the structure and movement of neuronal filopodia.
Ryan Hylton, Jessica Heebner and Michael Grillo of Penn State College of Medicine also contributed to this research.
More information: Ryan K. Hylton et al, Cofilactin filaments regulate filopodial structure and dynamics in neuronal growth cones, Nature Communications (2022). DOI: 10.1038/s41467-022-30116-x
Journal information: Nature Communications
Provided by Pennsylvania State University