New software to help discover valuable compounds
As a postdoctoral research associate in the lab of BTI faculty member Frank Schroeder, Max Helf saw his labmates continually struggle when they were analyzing data. So, he decided to do something about it and developed a free, open-source app called Metaboseek, which is now essential to the lab's work.
The Schroeder lab studies the roundworm Caenorhabditis elegans, one of the most successful model systems for human biology, to discover new metabolites that govern evolutionarily conserved signaling pathways and could be useful as leads for the development of new pharmaceuticals or agrochemicals. The researchers accomplish this task by comparing the metabolites between two different worm populations—a process called comparative metabolomics.
Given that samples routinely have more than 100,000 compounds in them, computational approaches are essential to perform the analysis.
The team had been relying on software packages that did not offer the required level of flexibility to easily customize analysis parameters. That limitation, and the lack of a suitable graphical user interface, meant Helf's colleagues faced the cumbersome task of visually inspecting mounds of data—for example, to spot possible false positives—and jumping between several other software tools to confirm and filter out those meaningless results.
"It just seemed very inefficient to me, and I couldn't get over the shortcomings of other software solutions for this problem," Helf said. "I thought there had to be an easier way, so I started to write code for my own software."
Helf developed the initial version of his software in 2017, and continued to improve it over the next two years. "Besides addressing the problems my labmates were already facing, I talked to them about what else held them back—what they wanted to do but weren't even trying—and built those features in the app," said Helf, who is now a bioinformatics product manager at proteomics company Biognosys AG. "I wanted this new tool to be user-friendly and accessible to anyone who does chemical biology."
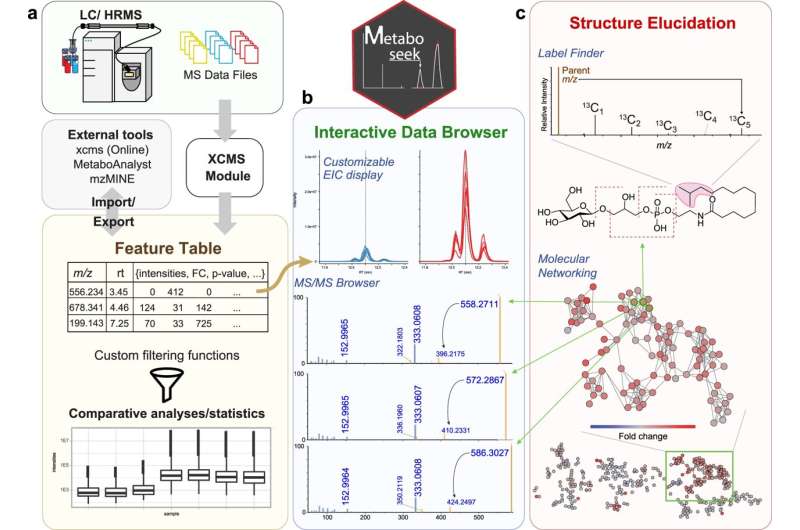
The result was Metaboseek, an app with a graphical interface that incorporates multiple data analysis tools that non-coding researchers would otherwise not have. The app streamlines the analysis of comparative metabolomics data by helping the researcher determine which data features are real and letting them dig deeper into those features—all within the same tool.
"Max did this without me even requesting it," Schroeder said. "Before I knew that this was happening, there was Metaboseek. We started using it, and now our lab and many collaborators couldn't exist without it."
In a study published in Nature Communications on February 10, Schroeder's team provided proof-of-concept for Metaboseek by applying it to an important fat metabolism pathway that hadn't yet been studied: the α-oxidation pathway in C. elegans that helps break down a class of fatty acids.
Using Metaboseek, the team found that roundworms lacking a key gene in the α-oxidation pathway accumulated hundreds of previously unreported metabolites. The findings are important because α-oxidation is a basic biochemical pathway in worms that is conserved in humans, Schroeder said.
"Bennett Fox did the chemistry work, so this study was a nice collaboration between the two postdocs," added Schroeder, who is also professor in Cornell University's Department of Chemistry and Chemical Biology.
According to Schroeder and Helf, there are a few reasons why there aren't a lot of good analytic tools for comparing metabolomics data. First, comparative metabolomics is a relatively young field compared with other data-heavy fields of biology like genomics (which focuses on DNA) and proteomics (which focuses on proteins), so there hasn't been enough time to develop software tools and database infrastructure.
Additionally, over the last decade, the advent of affordable, ultra-high-resolution mass spectrometers for collecting metabolomics data has increased by perhaps more than tenfold the amount of data one sample can generate—creating an even greater need for sophisticated tools that can keep up with the flood of data.
Metaboseek meets these needs with an array of features for analyzing various types of data to aid compound identification, structure determination, assignment of metabolites to families based on structural similarities, tracking radiolabeled compounds, and more.
More information: Maximilian J. Helf et al, Comparative metabolomics with Metaboseek reveals functions of a conserved fat metabolism pathway in C. elegans, Nature Communications (2022). DOI: 10.1038/s41467-022-28391-9
Journal information: Nature Communications
Provided by Boyce Thompson Institute