Scientists pack centuries of organic chemistry into neat 2D visualization

Researchers from Skoltech, Lomonosov Moscow State University, and Sirius University of Science and Technology have proposed a new method for visualizing chemical reactions to help scientists understand the global chemical reaction space and come up with ways of synthesizing organic compounds used in the industry. Reported in ACS Omega, the neural network-based method projects chemical reactions onto a 2D plane as dots, grouping similar chemical reactions together.
Chemists are constantly on the lookout for new ways of synthesizing useful organic compounds. These may range from the active ingredients in drugs and pesticides to fuel additives and other industrially significant substances: organic LEDs, dyes and pigments, etc. Since there are many ways to synthesize an organic compound, medicinal chemists have to dig into large reaction databases. Even for a simple compound, one can find hundreds of already known synthetic ways. It is challenging to analyze this amount of data using only human perception.
"Analyzing a typical database search output, a chemist can group reactions of a similar kind together to get an idea of the compound's synthetic landscape, but this requires a well-established chemical intuition, and it may be subjective, too," Sergey Sosnin of Skoltech says.
To simplify this process and make it more consistent, the researchers devised a way to capture the "essence" of chemical reactions and plot them on a graph for easy analysis. "It is more convenient to look at a picture rather than a long list of reactions. We visualize reactions based on what the reactants and the products are," Sosnin adds.
The proposed method converts a molecule into a numerical representation (bit vector). Then the algorithm extracts the essence of the reaction by subtracting the vectors of the reagents from those of the products. "In a way, the resulting vector stands for whatever's changed in the reaction, regardless of which specific compounds were involved," Sosnin explains. "That's what makes it such a powerful and pure representation of a reaction."
The problem with reaction vectors is they are in themselves unintelligible—unless you're good at thinking in 1,024 dimensions.
"We visualize these vectors, which are inaccessible to direct human comprehension, using an approach known as—here comes a mouthful—parameterized t-distributed stochastic neighborhood embedding," the researcher comments. "A neural network projects each multidimensional vector to the coordinates of a point on a plane."
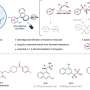
Given this chart, a chemist can recognize typical reaction types, for example the clusters indicated by diamonds numbered one through three in figure 1. Suppose someone is interested in ways of synthesizing the anti-HIV/AIDS drug darunavir (purple circles) or asthma medication montelukast (gray circles). The visualization affords insights as to which reaction types are mostly used for the purpose, which appear underused—or perhaps not used at all—despite possible assumptions to the contrary on the part of the researcher.
The team stresses the objective nature of the visualization. This is a bit like classifying animals based on DNA only, without ever having taken a single look at them. You may find, for example, that falcons, surprisingly, are more closely related to parrots than to other birds of prey. With chemical reactions, faulty intuitions can play similar tricks on us.
More information: Mikhail Andronov et al, Exploring Chemical Reaction Space with Reaction Difference Fingerprints and Parametric t-SNE, ACS Omega (2021). DOI: 10.1021/acsomega.1c04778
Journal information: ACS Omega
Provided by Skolkovo Institute of Science and Technology