An unexpected attraction between nucleic acids and fat
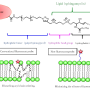
All cells on Earth are enveloped in a fatty layer of lipids. Lipid membranes protect the content of the cells, including genetic information such as RNA and DNA. A new study from the researchers at the B CUBE—Center for Molecular Bioengineering at TU Dresden reveals how lipids and RNA can directly interact and how this regulates RNA activity in unexpected ways. The study could help explain how RNA could be regulated in primordial or synthetic biological systems and lead to improvements in the design of RNA vaccines.
Lipids and RNA are two of the essential building blocks of cells. Ribonucleic acid (RNA) is a versatile molecule. On the one hand, like DNA, RNA encodes genetic information which is essential for life to replicate itself. On the other hand, like proteins, RNA molecules carry out chemical reactions. One molecule that can carry the information and also execute its instructions through chemical reactions is an attractive feature to synthetic biologists who try to build minimal living systems from scratch. However, finding a simple way to control RNA activity has always been a key challenge.
In a new study published in the PNAS journal, Tomasz Czerniak and James Sáenz discovered that lipids can directly modulate the activity of RNA in a simple synthetic system. "This opens up a completely new path to think about how we can use RNA-lipid interactions for bioengineering, for example, delivery of mRNA therapies," says Dr. James Sáenz, research group leader at the B CUBE—Center for Molecular Bioengineering and the senior author of the study. In addition, their work can also help to give clues about the origin of ancient life. A popular hypothesis on the origin of life is that life itself likely emerged from self-replicating RNA molecules, years before the evolution of DNA and proteins. Here, the simple and effective way of modulating RNA activity would be the key for the organization of early life on an ancient Earth.
The RNA-lipid world
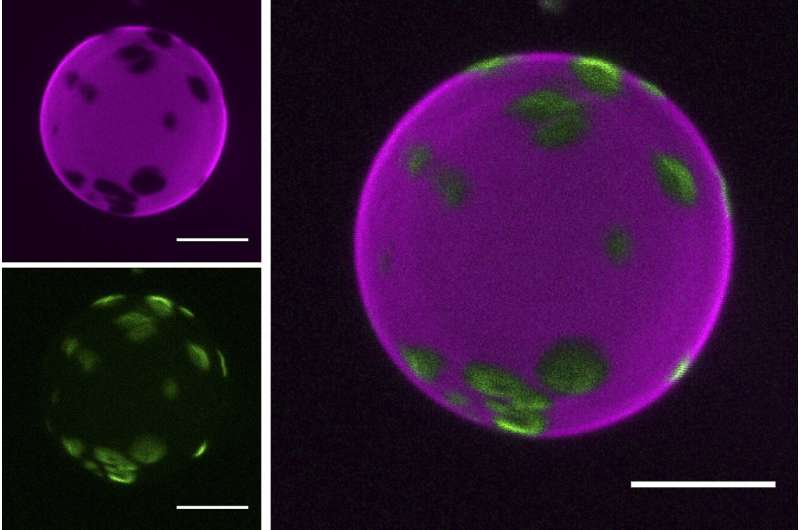
The scientists tested how well different types of RNA molecules interacted with lipid membranes. They found that some RNAs were binding lipids better than others and that this depended on the sequence and structure of the RNA molecule. In particular, guanine—one of the four building blocks that makes up RNA, was crucial for RNA to stick to the lipids. Adding additional chains of guanines to the RNAs made them even stickier, providing a way to control the strength of RNA-lipid interactions.

It turned out that guanine not only directly enhanced RNA-lipid binding but also made it stickier by promoting the folding of RNAs into different structures. One such structure, called a G-quadruplex, is found in cells and is known as an important element of RNA activity and regulation. "This raises the possibility that RNA-lipid interactions could be still happening in modern cells, possibly as a remnant of a long-extinct RNA-lipid world," says Sáenz.
Lipids give RNA self-control
Once the researchers had discovered how to engineer RNA to stick better to lipids, they then showed that such RNA-lipid interactions could be used to control the activity of RNAs that catalyze chemical reactions. "It's the first time that I'm aware of, where someone has demonstrated sequence-specific effects in the way a lipid can influence RNA catalysis," says Professor Gerald Joyce of the Salk Institute for Biological Studies in California, who was not involved in the study.
The future of RNA-lipid interactions
"As a next step we would like to understand how to use RNA-lipid interactions to engineer synthetic life and if these interactions are important in modern organisms, including humans," explains Sáenz. The team also points out that the insights from their work could provide new ways of designing lipid nanoparticles for mRNA vaccine formulations.
More information: Tomasz Czerniak et al, Lipid membranes modulate the activity of RNA through sequence-dependent interactions, Proceedings of the National Academy of Sciences (2022). DOI: 10.1073/pnas.2119235119
Journal information: Proceedings of the National Academy of Sciences
Provided by Dresden University of Technology