Researchers use CARC resources to explain genetic histories of island birds
University of New Mexico Ph.D. candidate Ethan Gyllenhaal has been engaged in a series of research projects that use genomic analyses to explain the complex evolutionary processes that take place within and between island bird populations. He is conducting this research in collaboration with researchers from the UNM Andersen Lab and the University of Kansas Moyle Lab.
One of his recent projects, outlined in a paper published in Molecular Ecology, explains the existence of an unexpected genetic finding in Australian Spectacled Monarch birds.
Most of us think that animals fit neatly into one discrete, static category called a species, but the reality of animal classification is far more complex. Sometimes, one species can separate into two when groups of animals are geographically separated and allowed to adapt to different environments.
This phenomenon, called allopatric speciation, is responsible for much of the biodiversity around us, like the stark difference, for instance, between squirrels on either side of the Grand Canyon. Other times, species can blend into each other to form a continuum, as Mallet, Beltrán, Neukirchen, and Linares observed in the Heliconius genus of butterflies found throughout the Americas.
Gyllenhaal's work focuses on how these types of speciation processes happen in bird populations living on and around islands. One of his recent projects used genomic analysis to uncover the history of interbreeding between the Spectacled Monarch birds of Australia and the Spot-Winged Monarch birds of nearby New Guinea.
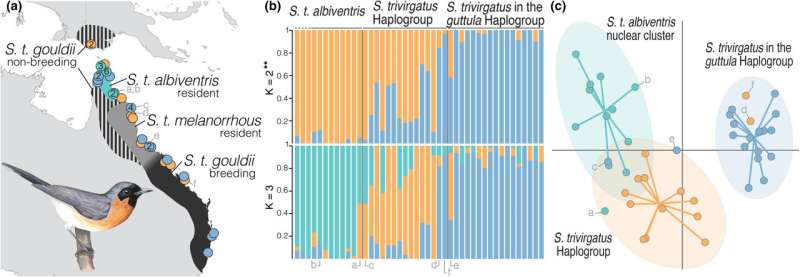
Analysis of Spectacled Monarch samples had shown a curious genetic finding: the mitochondrial DNA (the specialized DNA used by the power-generating mitochondria of a cell) of some Spectacled Monarch specimens did not match the nuclear DNA that is used by entire cells. These Spectacled Monarch samples appeared to contain the mitochondrial DNA of Spot-Winged Monarchs, leading to questions about a possible history of interbreeding between the two species.
To explain how this genetic rarity occurred, the Andersen Lab team needed to create a consensus reference assembly for the targeted genomic regions—a DNA sequence code that can be used to represent the DNA of all members of a population—for the Spectacled Monarch and related species. This type of reference requires an incredible quantity of computational memory to assemble, so Gyllenhaal and his colleagues used the Xena high-memory nodes at the Center for Advanced Research Computing to perform their computations.
Although it is nearly impossible to conclude with certainty that natural selection had no part in the development of genetic discordance in Spectacled Monarchs, the Andersen group's findings are consistent with the conclusion that crossbreeding took place, long ago, between the Australian Spectacled Monarch and the New Guinean Spot-Winged Monarch.
Gyllenhaal explains, "I think the Australia-New Guinea divide was somewhere in the realm of, I believe, 2-3 million years [ago]… and the mitochondrial DNA seemed to be, as we call it, 'captured' from the New Guinea population into the Australian population somewhere in the order of… 200 to 400 thousand years ago."
In other words, at some point around 200–400,000 thousand years ago, the geography of the region allowed New Guinean birds to crossbreed with Australian birds, leaving some Spectacled Monarchs with Spot-Winged Monarch mitochondrial DNA that persists in the population today. This was due to the lower sea level during the Pleistocene glacial cycles, which exposed a vast expanse of land connecting the two land masses, known as the Sahul shelf.
Since the publication of this paper, Gyllenhaal has continued his study of island birds with four different projects. His ongoing research examines bird species distinctions in the Solomon Islands, bird plumage variation in Fiji, and the island biogeographic factors that affect evolutionary changes in birds.
Gyllenhaal summarizes the findings of his research to date, reporting, "[One of] the two main takeaways from both the published and the ongoing work is that on islands, you get a lot of things that seem like they're going to be [discrete] species, but when they have that ability to come back into contact, they will exchange genes readily to the point of populations fusing. The other one is just… the porous[ness of the] boundaries between these species that live on different islands and have never essentially had their species boundaries tested."
More information: Michael J. Andersen et al, Complex histories of gene flow and a mitochondrial capture event in a nonsister pair of birds, Molecular Ecology (2021). DOI: 10.1111/mec.15856
Journal information: Molecular Ecology
Provided by University of New Mexico