Ancient marsupial 'junk DNA' might be useful after all
Fossils of ancient viruses are preserved in the genomes of all animals, including humans, and have long been regarded as junk DNA. But are they truly junk, or do they actually serve a useful purpose?
Researchers at UNSW Sydney, who examined the DNA and RNA of 13 Australian marsupial species, believe that viral fossils may be helping protect animals from infection.
"These viral fragments have been retained for a reason. Over millions of years of evolution, we would expect all DNA to change, however these fossils are preserved and kept intact," says doctoral student and lead researcher Emma Harding, from UNSW's School of Biotechnology and Biomolecular Sciences.
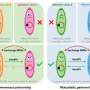
"Our study shows how the viruses buried in the animals' DNA are used to make what we call non-coding RNAs, which carry out tasks inside the animal cells such as protecting against outside infection.
"The animal DNA has basically grabbed a viral sequence—which used to harm it—and ends up using it for its own benefit."
If Ms Harding is right, her theory may have important implications for our understanding of the role of viral fossils in our own DNA and the animal kingdom in general. For example, viral fossils can be turned into RNA that binds specifically to the virus—and destroys it—if it tries to infect the cell again.
"This could be a mechanism similar to vaccination, but is inherited through generations. By keeping a viral fossil, the cell is immunized against future infection.
"If we can show it occurring in marsupials, it may also be occurring in other animals, including humans.
"So if we look more closely at the viral fossils inside our DNA, we could potentially get clues about how they may be protecting us," Ms Harding says.
Evolutionary rewind
To understand how Ms Harding arrived at her conclusions, it's worth stepping back a hundred million years to a time when Australia was once part of the Gondwanaland super-continent along with South America, Africa and Antarctica.
Scientists think the first marsupials originated in South America and later migrated to Australia via Antarctica. The lack of competing mammals in these southern regions may have allowed the marsupials to further evolve, with about 250 species now living in Australia, and about 120 in South America.
Ms Harding says when we look at the genetic makeup of Australian marsupials, the presence of viral fossils, known as endogenous viral elements (EVEs), are like time stamps marking when an animal was infected.
"My research looks at EVEs in Australian marsupials to firstly identify what types of viruses have integrated, and secondly investigate if they play an active role in the marsupial cells.
"One of the EVEs I found was from the Bornaviridae family of viruses, which first entered the animals' DNA during the time of the dinosaurs when the South American and Australian landmasses were still joined together.
"Bornaviridae viruses were previously thought to have evolved 100 million years ago. But the one I found in almost every marsupial DNA we looked at puts it at 160 million years old—so it was great to be able to contribute to our knowledge of evolutionary history.
"By understanding how viruses have evolved, we can predict how they may change in the future, and plan strategies to prevent outbreaks."
The work
The researchers examined multiple transcriptomes—the collection of all RNA in a cell—of Tasmanian devils, tammar wallabies, long-nosed bandicoots, fat-tailed dunnarts, bare-nosed wombats, koalas and sugar gliders. They also looked at single transcriptomes of a false antechinus, Southern brown bandicoot, striped possum, Western pygmy possum, brushtail possum and a yellow-footed rock wallaby.
They used the genetic sequence of modern viruses, such as Ebola and Zika, to hunt through the transcriptomes and look for matches—fragments of ancient marsupial viruses. The benefit of searching through the transcriptome rather than the genome is that it finds viral fragments that are still 'switched on' and transcribed—making RNA in the cells of marsupials, as opposed to stagnant fossils.
"Transcription is evidence that the EVEs are active in cells and possibly doing something functional," says Ms Harding.
Out of more than 130 known viral families, three were seen again and again throughout the marsupial species.
Bornaviridae, Filoviridae and Parvoviridae are commonly seen in eutherian (or placental) mammals, including humans. Ms Harding says she noticed a unique trend in marsupials where different parts of the viral genomes were retained as fossils over time.
"It was interesting to see that the DNA blueprints for two viral proteins—the nucleocapsid, which is the shell of the virus, as well an enzyme essential for replication—were very common in the marsupial.
"Our hypothesis is that these two proteins are good targets for antiviral defense, therefore copies of them have been kept in the genome. These copies can block areas of the incoming viral genome so the virus cannot properly make these proteins. Without the proteins, it cannot replicate and the immune system can clear the infection."
Protected species
The researchers also found evidence for small RNA molecules which are known to target and break down foreign viral RNA strands, adding more weight to the virus-fighting hypothesis.
"I found that some EVEs were spliced from the genome to create siRNA and piRNA—two types of molecules that are used in the immune system of plants and animals including humans," Ms Harding says.
"These RNA molecules were enriched in the testis of the male marsupials, and may be helping to protect future offspring from viral infections. Marsupials have a very different development to other mammals, and are born without a fully functional immune system, so these small molecules may help them defend them while they are still growing."
The study was based on the available RNA sequence data of 35 samples from 13 marsupials. Ms Harding hopes future laboratory research will prove their role in viral immunity, which could lead to the development of novel RNA antivirals against a large range of viruses.
The research was published recently in Virus Evolution and a review is published in Microbiology Australia.
More information: Emma F Harding et al, Ancient Viral Integrations in Marsupials: A Potential Antiviral Defence, Virus Evolution (2021). DOI: 10.1093/ve/veab076
Provided by University of New South Wales




















