New technique to manipulate gene expression and study genetic diseases
Emma Andersson, senior researcher at the department of Cell and Molecular Biology, and Karin Mangold, Ph.D. student, have recently published an article in Cell Reports Methods in which they developed a new technique to reduce the use of mice and to get faster results.
Understanding how our genes control how we develop from a fertilized egg to a fully functional human being requires experiments in which we "take away" or "add to" parts of the genes (pieces of DNA) to see what they are needed for, or what they can do. Often, scientists use mice to understand how genes control development and disease in mammals, but taking away and adding genes in mouse models has previously been slow work (months to years), that could also unfortunately lead to the production of mice that could not be used. To solve this problem, Emma Andersson and her group at the Department of Cell and Molecular Biology developed a method to quickly take away or add genes in mice (days/weeks), focused on the brain and spinal cord.
"We showed that the method was very efficient, and could be used to understand the functions of genes associated with human disease. With this new technique, it will be faster and easier to study genes in mice, so we can test how different genes (discovered to be mutated in different patients) actually control health and disease." Emma Andersson explains.
The technique can also be used to systematically and quickly test a lot of different genes (screening), or for "lineage tracing"—understanding what each stem cell in the nervous system can give rise to, by labeling these and following the cells over time.
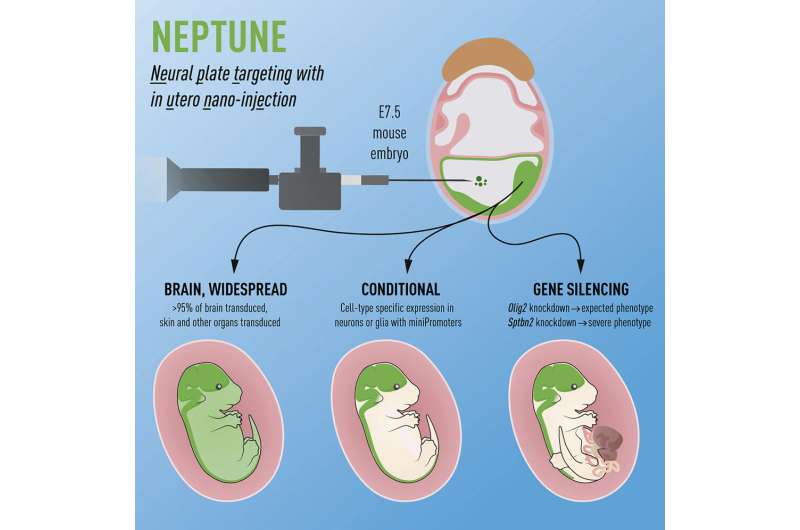
The technique is conceptually very similar to amniocentesis which is done on pregnant women to test for genetic disease in the unborn child. The researchers inject a DNA-modifying virus into the amniotic fluid of sedated pregnant mice, which can then modify the cells in contact with the amniotic fluid. They technique doesn't touch the embryos themselves, and the female mice wake up after a few minutes and are running around and grooming themselves normally.
Emma and her group named the technique "NEPTUNE" for "NEural Plate Targeting with in Utero Nano-injEction," in part as a reference to Neptune the god of water (and life-giving amniotic fluid) and in part as a reference to the blue planet in our solar system, since the uterus in pregnant mice (who can carry up to 20 embryos in one pregnancy) can be seen as a series of beads (embryos) on a string (along the uterus), which also makes us think of planets in the Solar system. All the worlds' potential in embryos and because it was an easy name to remember.
"We can now use NEPTUNE to answer many unresolved questions in developmental neuroscience. We are also developing the technique to be able to manipulate other organ systems during development. This can reduce the usage of mice in life science by over 90-95%. We have established a core facility so that others have easy access to the technique (INFINIGENE), and we hope that we can thus contribute to reducing the numbers of animals used in research. All in all, we plan to use this exciting technique to understand how stem cells develop in embryos, which genes are most important at specific stages, and why some genes are associated with genetic disorders, especially in children."
More information: Katrin Mangold et al, Highly efficient manipulation of nervous system gene expression with NEPTUNE, Cell Reports Methods (2021). DOI: 10.1016/j.crmeth.2021.100043
Provided by Karolinska Institutet