Method for discovery of antiviral drugs
The current COVID-19 pandemic has highlighted the need for methods to identify new or repurposed drugs as antivirals. Researchers at Uppsala University and Karolinska Institutet are now presenting a new screening approach that focuses on the identification of virus-specific morphological changes in virus-infected cells.
The new method focuses on identifying the changes (morphological profiles) that the virus induces on the infected cells by using a modified version of the Cell Painting protocol, an established assay that uses a cocktail of fluorescent reagents to stain several cellular compartments. These morphological profiles are then used as a basis to screen for drugs that can reverse the virus-induced effects.
In a single assay that combines Cell Painting with antibody-based detection of viral infection at a single cell level, the researchers have not only been able to confirm the antiviral effect of known reference drugs, but also to identify novel compounds as potential antivirals. The method includes image and data analysis pipelines using CellProfiler, a popular image analysis software, which the researchers have made openly available to facilitate the use and spread of this new method.
The majority of the methods for the discovery of antiviral drugs that are available today tend to focus on the effects of such drugs on a given virus, its constituent proteins, or enzymatic activity. However, the consequences for host cells are often neglected.
"This is a problem, as potential toxicity impacting the overall physiology of host cells may mask the effects of both viral infection and drug candidates. With our method, on the other hand, we are able to assess the general health of host cells, and in parallel identify antiviral properties of compounds" says Jordi Carreras-Puigvert, senior author of this work and lecturer at the Pharmaceutical Bioinformatics group, Department of Pharmaceutical Biosciences at Uppsala University.
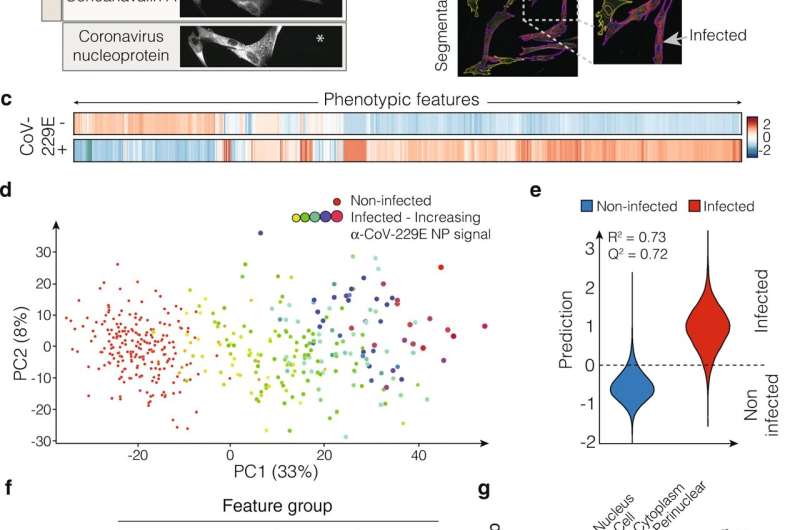
"The reasoning behind our approach was to obtain unbiased morphological profiles in the context of the host cell to study viral infection and compound treatment in a single assay. We modified the Cell Painting protocol, combining it with a virus-specific antibody staining. This enabled us to select the virus-infected cells with high precision and even relate the morphological profiles with the viral protein levels in each cell," says Jonne Rietdijk, first author of this work and Ph.D. student at the Pharmaceutical Bioinformatics group, Department of Pharmaceutical Biosciences at Uppsala University.
The researchers show that their method can successfully capture virus-induced phenotypic signatures of human lung fibroblasts infected with human coronavirus. They also demonstrate that the method can be used in phenotypic drug screening using a panel of nine host- and virus-targeting antivirals, and that treatment with effective antiviral compounds reversed the morphological profile of the host cells towards a non-infected state.
"By only doing minor adjustments to the image analysis pipeline that we provide, we anticipate that our untargeted approach, will enable other applications using diverse (human-derived) cell lines, as well as different viruses," says Jonne Rietdijk.
"There are two main uses that we see for the method, one is the screening for already medically available drugs that could be repurposed as antivirals. The second one is the screening for actual novel compounds. Since we create compound-specific signatures, we can then compare these to a set of signatures extracted from compounds with known mode of action, thereby potentially identifying the target of a given novel compound, in this case an antiviral," says Jordi Carreras-Puigvert.
More information: Jonne Rietdijk et al, A phenomics approach for antiviral drug discovery, BMC Biology (2021). DOI: 10.1186/s12915-021-01086-1
Journal information: BMC Biology
Provided by Uppsala University