Plants get a faster start to their day than we think
To describe something as slow and boring we say it's "like watching grass grow", but scientists studying the early morning activity of plants have found they make a rapid start to their day—within minutes of dawn.
Just as sunrise stimulates the dawn chorus of birds, so too does sunrise stimulate a dawn burst of activity in plants.
Early morning is an important time for plants. The arrival of light at the start of the day plays a vital role in coordinating growth processes in plants and is the major cue that keeps the inner clock of plants in rhythm with day-night cycles.
This inner circadian clock helps plants prepare for the day such as when to make the best use of sunlight, the best time to open flowers for pollinators and release pollen and when to get ready to respond to drought conditions.
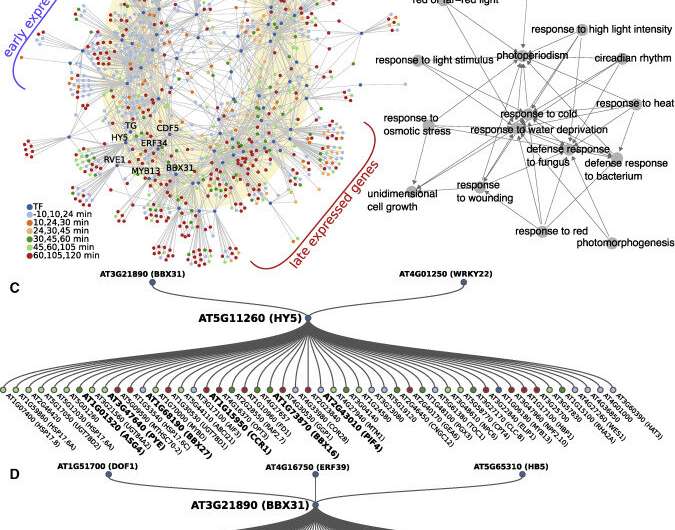
There is a peak of gene activity within an hour of dawn; many of these genes code for transcription factors—proteins that regulate expression of a host of downstream genes—with roles related to light, stress and growth hormones, but the detail of how this peak is controlled is not understood.
Researchers at the Sainsbury Laboratory Cambridge University (SLCU) and University of York set out to investigate this burst of activity so as to better understand what happens at the genetic level by sampling thale cress, Arabidopsis thaliana, every two minutes from dawn to measure gene activity.
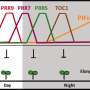
"We set out to characterize 'dawn burst' dynamics in more detail, focussing on the expression of transcription factor genes. We found three distinct gene expression waves within two hours after dawn. The first of these occurs just 16 minutes after dawn and lasts only 8 minutes." said Dr. Martin Balcerowicz, researcher at SLCU and first author of the research published in Molecular Plant.
"Many of these genes are known to be sensitive to light and temperature, but we wanted to find out specifically how the transcription of these genes is coordinated. Interfering in photoreceptor signaling, the circadian clock and chloroplast derived light signals did cause problems in some genes' expression, but there was a large proportion of genes still unaffected. This indicated to us that some of the upstream pathways are redundant and that additional regulators are in play."
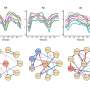
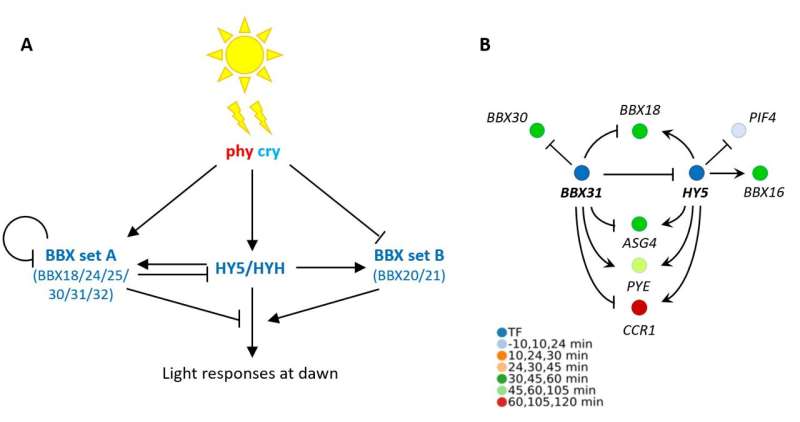
The team integrated their data with already published transcription factor-DNA binding data and identified a gene regulatory network at dawn, with two key regulators of light signaling—HY5 and BBX31—at its core. These transcription factors are known to jointly control de-etiolation, which is the developmental switch a seedling undergoes when it emerges from the soil, experiencing light for the first time, and starts greening and unfolding its leaves. It appears that these genes also play a central role during the dark-to-light transition at dawn.
"In fact, multiple BBX genes form part of the dawn burst alongside BBX31, HY5 and its homologue HYH," says Dr. Balcerowicz. "These genes include both positive and negative regulators of the light response. We found that they act downstream of phytochrome and cryptochrome photoreceptors to control a light-induced subset of dawn burst genes, with HY5 and BBX31 having largely antagonistic roles. This observation strengthens the idea that HY5 and BBX genes act in concert to fine-tune light responses in the context of the day-night cycle."
Dr. Daphne Ezer, lecturer in Computational Biology at the University of York and senior author of the study, investigates gene-environment interactions through analysis of gene networks. "By studying gene networks we can interpret how plants integrate light and temperature signals in the early morning to entrain the circadian clock. Taken together, our results show that phytochrome and cryptochrome signaling is required for fine-tuning the dawn transcriptional response to light, but separate pathways can robustly activate much of the program in their absence."
"Characterizing the peak we see in gene expression that results from the onset of light is useful in helping us to understand how plants respond to light and, in particular, for crops grown under artificial lighting, how this dawn burst impacts longer term on growth."
More information: Martin Balcerowicz et al, An early-morning gene network controlled by phytochromes and cryptochromes regulates photomorphogenesis pathways in Arabidopsis, Molecular Plant (2021). DOI: 10.1016/j.molp.2021.03.019
Journal information: Molecular Plant
Provided by University of Cambridge