Watch me move it, move it: Gliding structure in Mycoplasma revealed
Much of human invention and innovation has been the result of our discovery and replication of natural phenomena, from birds in flight to whales that dive deep into the ocean. For the first time, researchers have captured at the nanometer level the gliding machinery of the bacterium Mycoplasma mobile. Their findings were published in mBio. It illuminates the origin and operating principle of motility, which could serve as a basis for the next generation of nanoscale devices and pharmaceuticals.
"My lab has been studying the molecular nature of bacteria from the Mycoplasma genus for years," states Professor Makoto Miyata from the Graduate School of Science, Osaka City University and lead of the research group. "And we have developed a conceptualization of how some of these parasitic bacteria 'glide' around their hosts."
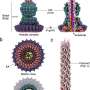
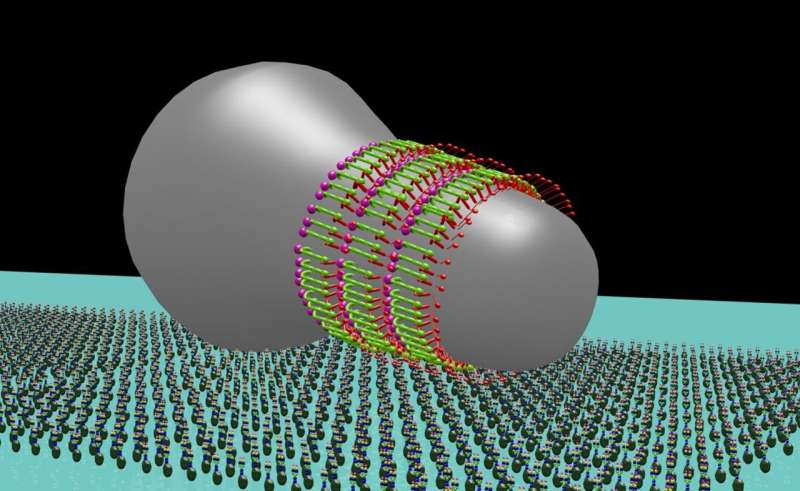
For example, Mycoplasma mobile forms a protrusion at one end giving the bacterium a flask shape. At the tapered end are external appendages that bind to solid surfaces, and in concert with an internal mechanism, cause the bacterium to glide across the surface of its host to find nutrient-rich sites and escape the host's immune response.
"What we lacked was a visual understanding of the internal mechanism," says first author Kohei Kobayashi, "and for this, we needed the right technology." In collaboration with a research team led by Professor Noriyuki Kodera and Professor Toshio Ando of Kanazawa University, Prof. Miyata and his team used high-speed atomic force microscopy, a cutting-edge technique that can visualize biological molecules in action at nanometer and sub-second spatiotemporal resolution. They scanned M. mobile cells from the outside and successfully visualized the internal structural movement in real time.
First, the team immobilized live M. mobile on a glass substrate and probed the cell surface with the fine needle of HS-AFM, confirming the structure according to past measurements taken with electron microscopy. Then the team visually differentiated the internal structure from the external appendages by computationally extracting the signals hidden in the video images. What they discovered was an internal chain structure causing the external appendage structure to move 9 nanometers right, relative to the gliding direction, and 2 nanometers into the cell interior in 330 milliseconds and then return to their original position, based on ATP hydrolysis.
"In the future, we intend to isolate the molecular motors and analyze the cells with higher spatial and temporal resolution, and through electron microscopy, understand the mechanism for the gliding motion at the atomic level," says Prof. Miyata. An atomic understanding of this most complicated mechanism of motility may be the key to human replications of it.
More information: Miyuki S. Nishikawa et al, Refined Mechanism of Mycoplasma mobile Gliding Based on Structure, ATPase Activity, and Sialic Acid Binding of Machinery, mBio (2020). DOI: 10.1128/mBio.02846-19
Journal information: mBio
Provided by Osaka City University