The atomic makeup of M. pneumoniae's "Nap" protein complex glides into view

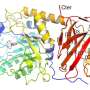
Using X-ray crystallography and cryo-electron microscopy, an international team of scientists unravel the atomic structure of the proteins P1 and P40/P90 which make up the "Nap" structure—a protein complex that the bacterium M. pneumoniae uses to attach and move around human cells to cause pneumonia. This will allow us to better understand the "Nap" structure and develop medicine and vaccines that stop the bacterium from infecting humans.
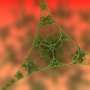
Caterpillars are small bugs with two rows of tiny legs that all move in a coordinated manner, causing the caterpillar to almost glide across a leaf. Mycoplasma pneumoniae, the bacterium behind many cases of pneumonia in human communities, moves in a similar fashion around our body's cells.
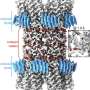
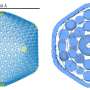
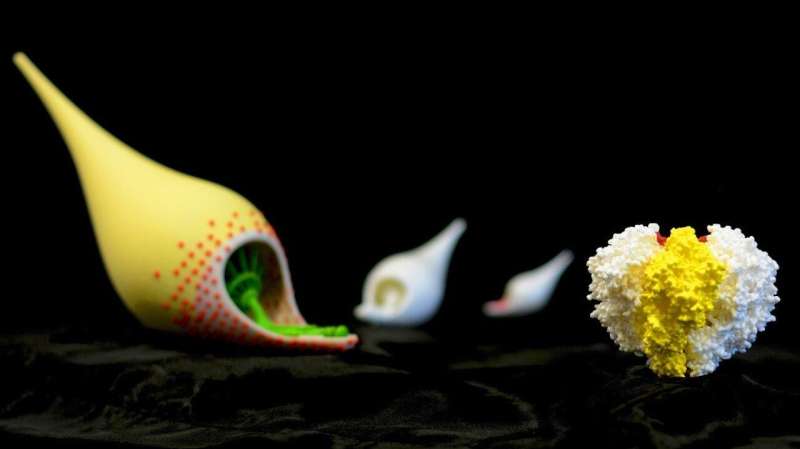
Using a lock and key attachment method, the yellow bowling pin shaped M. pneumonia (see left-hand object in the image) uses tiny microscopic appendages (the red dots) to attach itself to the surface of a cell and move around, altering itself to avoid detection from our immune system. In the 80's, scientists discovered a protein complex at the end of this appendage to be at the center of M. pneumoniae's ability to attach, move, and change. Called "Nap," this protein complex acts as a "key" that locks onto a sugary substance found all over the surface of our cells made up of sialic acid oligosaccharides. Understanding this "Nap" structure could lead to treatments that inhibit this locking mechanism, allowing for the production of medicine and vaccines that could put an end to the bacterium's ability to attach, move, change, and ultimately infect.
However, its unraveling had eluded the scientific community for more than 40 years—until now, when a team of researchers from 3 countries, including Japanese teams led by Professor Makoto Miyata of the Graduate School of Science, Osaka City University, clarified it at the atomic level.
The results of the research have been published in the October edition of the online scientific journal Nature Communications.
By isolating the proteins P1 and P40/P90 that make up the "Nap" structure and examining each one through a combination of X-ray crystallography and cryo-electron microscopy, "we found that the shape of the structure resembles four corndogs bundled at the stick and stuck into the membrane of M. pneumonia," explains Prof. Miyata. (the "Nap" structure is on the right-hand side of the image.) "Also, contrary to popular belief, we found it was the proteins P40/P90, not P1, that bind to the sialic acid substance found on our cell tissue."
"This is one of many big revelations to come, including how the bacterium avoids our immune system, now that the architecture of the "Nap" protein complex has been revealed," exclaims the professor.
More information: David Vizarraga et al. Immunodominant proteins P1 and P40/P90 from human pathogen Mycoplasma pneumoniae, Nature Communications (2020). DOI: 10.1038/s41467-020-18777-y
Journal information: Nature Communications
Provided by Osaka City University