New method of artificially creating genetic switches for yeast
A group of researchers from Kobe University and Chiba University has successfully developed a flexible and simple method of artificially producing genetic switches for yeast, a model eukaryotic organism. The group consisted of Researcher Tominaga Masahiro, Associate Professor Ishii Jun and Professor Kondo Akihiko (of Kobe University's Graduate School of Science, Technology and Innovation/Engineering Biology Research Center), and Professor Umeno Daisuke et al. (of Chiba University's Graduate School of Engineering).
Genetic switches are gene regulatory networks that control gene expression. The researchers established a platform for creating genetic switches that could be applied to the development of sophisticated, artificially controlled yeast cells to produce large quantities of valuable compounds. These research results were published in Nature Communications.
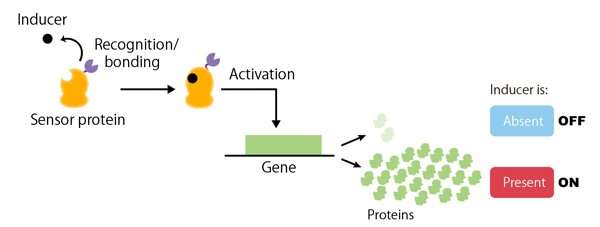
The number and type of genes that an organism possesses do not solely determine its life functions. The timing and quantity of proteins produced by a gene (i.e. gene expression) are other factors that are known to result in significant alterations. In the field of synthetic biology, recent advances have made it possible to generate many novel cell functions by artificially controlling the expression of certain genes. Genetic switches are necessary in order to control the rate and timing of gene expression. A genetic switch (Figure 1) is a regulatory system that turns the expression of a particular gene 'on' or 'off' in response to a stimulus (or inducer) from either inside or outside the cell (for example, the presence of a chemical substance). Consequently, genetic switches are an essential tool for synthetic biology, which aims to artificially design and construct cellular functions.
Many genetic switches have been developed for simple, single cell organisms (prokaryotes) such as E. coli. However, the systems of gene expression in eukaryotic organisms, such as humans, plants and yeast, are more complex. Consequently, there is a lag in the development of genetic switches for these organisms. Even though yeast is a model eukaryotic organism, attempts to engineer the functions of its cells have faced great limitations.
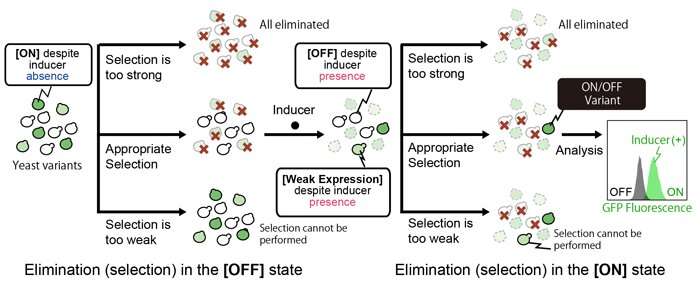
When constructing genetic switches, it is very difficult to predict where and how to alter the switches to enable gene expression to be controlled. Evolutionary molecular engineering is a useful method for determining this (Figure 2). The method involves creating a library of genetic switch variants by randomly inducing mutation in part of or the entire genetic switch, and then selecting the variants that show intended performance. Although it is easy to produce a large number of variants, the desired variants within this number must be quickly identified. An artificial process of elimination (selection) was carried out to select the cells that remained both when gene expression was 'off' and when gene expression was turned 'on' by a specific inducer. However, if the selection is too strong or too weak, it is not possible to single out the best variants. Although it is necessary to select functional genetic switch variants that are suitably robust in both 'on' and 'off' states, it is very difficult to predict how strong the selection should be beforehand.
The team of researchers from Kobe University and Chiba University established a workflow system whereby they could generate selections of varying strengths in parallel by changing the type or concentration of the chemicals used for selection. After selecting a group of variants, the researchers exposed each one to an external stimulus (inducer) and analyzed the extent to which this turned gene expression on by observing the change in the level of light emitted from GFPs (green fluorescent proteins). This allowed them to determine the most appropriate selection, in other words to easily identify genetic switch variants that demonstrated a high level of performance. Using this method, the researchers successfully developed three new genetic switches that were as efficient as the best-performing switch developed for yeast up until now.
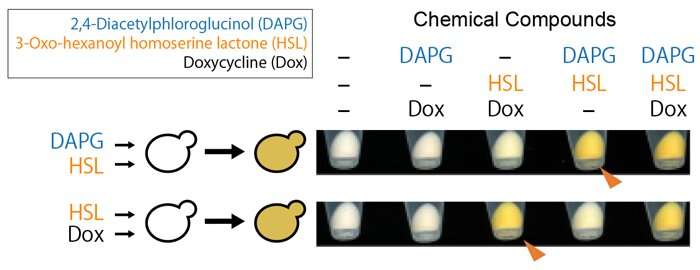
By integrating these three genetic switches, the researchers produced yeast that could biosynthesize orange pigment (β-carotene) under AND-gated control (i.e. where β-carotene could only be produced if two specific chemical compounds, DAPG and HSL, were present) (Figure 3).
The selection method developed by this research group will expedite the development of a wide range of genetic switches for yeast, with various performance levels and characteristics. This will also lead to a rapid increase in the number of genes that can be controlled in parallel. Combining these new genetic switches will make it possible to artificially design cellular functions. For example, this could contribute towards the development of sophisticated, artificially regulated yeast cells for producing large quantities of useful organic compounds.
More information: Masahiro Tominaga et al. Robust and flexible platform for directed evolution of yeast genetic switches, Nature Communications (2021). DOI: 10.1038/s41467-021-22134-y
Journal information: Nature Communications
Provided by Kobe University