How organisms filter out the noise to make accurate predictions

A new study by researchers from the University of Chicago and the French National Centre for Scientific Research shows how organisms filter information from their environment differently for a wide range of biological processes—from visually tracking the motion of objects to immune cells responding to pathogens—and then select the most useful inputs to respond accordingly.
Organisms constantly process information from the world around them to make predictions about the future. An animal like a larval salamander, for example, tracks the movements of small insects and crustaceans swimming around nearby. Using what it sees about how fast the prey is moving and in what direction it's heading, the salamander can gauge when and where it should strike for its next meal. But any biological system has constraints—the salamander can process only so much visual information at once, so it makes tradeoffs about which data are most important to help it catch its prey, and which data it can filter out.
Using this "information bottleneck" problem as a starting point, the researchers developed a series of equations that show how organisms weigh different variables and calculate sensory inputs to make predictions about the future most efficiently.
"All biological systems are constrained. They don't have infinite time, or infinite amounts of energy they can burn to do these calculations," said Assoc. Prof. Stephanie Palmer, a neuroscientist at UChicago and senior author of the study, which was published on March 8 in PLOS Computational Biology. "We know predictions happen at many different time scales in many different organisms, with many different mechanisms. So, what we're doing here is figuring out what the filtering or encoding should look like for a biological system trying its darndest to predict the future."
The study, led by UChicago biophysics graduate student Vedant Sachdeva, is a collaboration with biophysicists Thierry Mora and Aleksandra Walczak from the French National Centre for Scientific Research, the largest governmental research organization in France. The project grew out of an ongoing partnership between the two institutions to foster opportunities for research, education and scholarly engagement across a range of scientific fields. The agreement, the first of its kind between a U.S. university and the French National Centre for Scientific Research, funds other collaborations in such areas as molecular engineering, physics, computer science, mathematics, biochemistry, genetics, molecular biology and the social sciences.
Vedant and colleagues studied how the visual system and adaptive immune system filter information to make predictions under the information bottleneck framework. They first used the movements of an oscillating weight attached to a spring and bouncing around in a viscous medium, inspired by previous work by Palmer on how the salamander retina responds to a visual stimulus moving around in the same manner.
As expected, position and velocity are the most important variables to predict future movements, but how much each should be weighted depends on the movements themselves. The dampening in the system, i.e. the amount "bounce" the object has in response to random kicks and changes in direction, determines how important position is versus velocity. For example, in an overdamped system, where that bounciness is tightly constrained like a car with stiff shock absorbers, position is a more important predictor of future movements. In an underdamped system, like a car where the shocks are completely shot that bounces all over the place, both position and velocity are needed to predict future movements accurately.
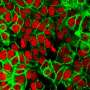
The researchers also considered other biological systems that rely less on short-term, adaptive processes like early vision does. The immune system has a long-term memory of sorts to keep track of pathogens. The adaptive immune system can make predictions about the genetic evolution of pathogens by comparing different versions it encounters, and projecting what future versions may look like to mount an appropriate response.
In this case, the immune system seems to favor preparing for a broad range of possible antigens versus trying to anticipate an exact match for the next virus. The team also looked at how historical information on the distribution and rate of genetic mutations can be used to predict evolution in a population over time.
Much of how an organism makes predictions in these different systems is hardwired evolutionarily, over time. The salamander knows a general set of rules for how different prey and predators in its environment tend to move. The immune system is adapted for certain types of pathogens that behave in predictable ways. But how do these systems adapt when conditions change in the real world?
Palmer said the team is working on that part of the problem next, taking what they've learned about how organisms make predictions and trying to understand how those systems adapt to new stimuli. In doing so, they hope to find some underlying rules that help explain predictions at a larger scale, across many different systems and time scales.
"We know prediction is broad and relevant in all these different contexts, and that the filters are a bit different depending on the problem," said Palmer, who is appointed in both the Department of Physics and the Department of Organismal Biology and Anatomy. "Now we can look at how they adapt in real time to changes, which I think will reveal things that are general and shared across these different systems."
More information: Vedant Sachdeva et al. Optimal prediction with resource constraints using the information bottleneck, PLOS Computational Biology (2021). DOI: 10.1371/journal.pcbi.1008743
Journal information: PLoS Computational Biology
Provided by University of Chicago