Supercomputers simulate new pathways for potential RNA virus treatment
University of New Hampshire (UNH) researchers recently used Comet at the San Diego Supercomputer Center at UC San Diego and Stampede2 at the Texas Advanced Computing Center to identify new inhibitor binding/unbinding pathways in an RNA-based virus. The findings could be beneficial in understanding how these inhibitors react and potentially help develop a new generation of drugs to target viruses with high death rates, such as HIV-1, Zika, Ebola, and SARS-CoV2, the virus that causes COVID-19.
"When we first started this research, we never anticipated that we'd be in the midst of a pandemic caused by an RNA virus," said Harish Vashisth, associate professor of chemical engineering at UNH. "As these types of viruses emerge, our findings will hopefully offer an enhanced understanding of how viral RNAs interact with inhibitors and be used to design better treatments."
Similar to how humans encode their genome using DNA, many viruses have a genetic makeup of RNA molecules. These RNA-based genomes contain potential sites where inhibitors can attach and deactivate the virus. Part of the challenge in drug development is that variations or mutations in the viral genome that may prevent the inhibitors from attaching.
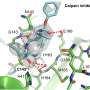
In their study, recently published in the Journal of Physical Chemistry Letters, Vashisth and his team created molecular dynamics simulations using the Comet and Stampede2 supercomputers to look specifically at an RNA fragment from the HIV-1 virus and its interaction with acetylpromazine, a small molecule that is known to interfere with the virus replication process.
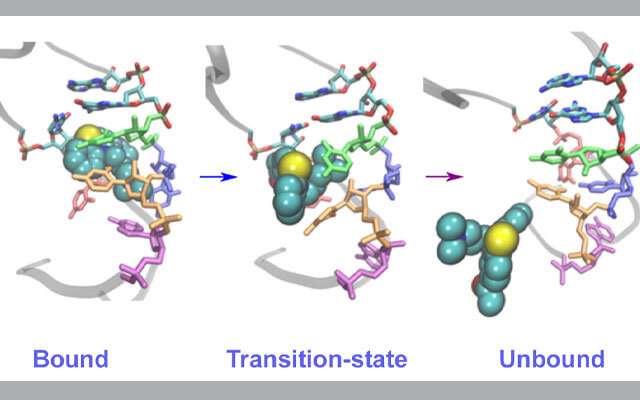
The scientists focused on the structural elements from the HIV-1 RNA genome because they are considered a good model for studying the same processes across a wide range of RNA viruses. These simulations enabled them to discover the pathways of the inhibitor unbinding from the viral RNA in several rare events—which are often difficult to observe experimentally—that unexpectedly showed a coordinated movement in many parts of the binding pocket that are the building blocks of RNA.
Thanks to the National Science Foundation's (NSF) Extreme Science and Engineering Environment (XSEDE) allocations on Comet and Stampede2, the researchers were able to run hundreds of simulations at the same time to observe what are called rare base-flipping events involved in the inhibitor binding/unbinding process that provided the new details of the underlying mechanism of this process.
"Our hope is that this adds new possibilities to a field traditionally focused on static biomolecular structures and leads to new medications," Vashisth said.
More information: Lev Levintov et al. Ligand Recognition in Viral RNA Necessitates Rare Conformational Transitions, The Journal of Physical Chemistry Letters (2020). DOI: 10.1021/acs.jpclett.0c01390
Journal information: Journal of Physical Chemistry Letters
Provided by University of California - San Diego