SMART designs tool to investigate bacteria behind hospital infections

Researchers from the Antimicrobial Resistance (AMR) Interdisciplinary Research Group (IRG) at Singapore-MIT Alliance for Research and Technology (SMART), MIT's research enterprise in Singapore, and Nanyang Technological University (NTU) have developed a tool using CRISPRi technology that can help understand and prevent biofilm development, drug resistance, and other physiological behaviors of bacteria such as Enterococcus faecalis.
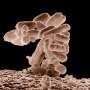
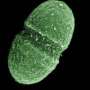
E. faecalis, a bacteria found in the human gut, is one of the most prevalent causes of hospital-associated infections and can lead to a variety of multidrug-resistant, life-threatening infections including bacteraemia (bloodstream infection), endocarditis (infection of the heart), catheter-associated urinary tract infection and wound infections.
However, current methods for understanding and preventing E. faecalis biofilm formation and development are labor-intensive and time-consuming. The SMART AMR research team designed an easily modifiable genetic technique that allows rapid and efficient silencing of bacteria genes to prevent infections.
In a paper titled "Multiplex CRISPRi System Enables the Study of Stage-Specific Biofilm Genetic Requirements in Enterococcus faecalis," published in the journal mBio, the researchers explain the scalable dual-vector nisin-inducible CRISPRi system which can identify genes that allow bacteria like E. faecalis to form biofilms, cause infections, acquire antibiotic resistance, and evade the host immune system. The team combined CRISPRi technology with rapid DNA assembly under controllable promoters, which enables rapid silencing of single or multiple genes, to investigate nearly any aspect of enterococcal biology.
"Infections caused by E. faecalis are usually antibiotic tolerant and more difficult to treat, rendering them a significant public health threat," says Dr. Irina Afonina, postdoctoral associate at SMART AMR and lead author of the paper. "Identifying the genes that are involved in these bacterial processes can help us discover new drug targets or propose antimicrobial strategies to effectively treat such infections and overcome antimicrobial resistance."
The team believes their new tool will be valuable in rapid and efficient investigation of a wide range of aspects of enterococcal biology and pathogenesis, host-bacterium interactions, and interspecies communication. The method can be scaled up to simultaneously silence multiple bacterial genes or perform full-genome studies.
"Bacterial biofilms are clusters of bacteria that are enclosed in a protective, self-produced matrix," says SMART AMR Principal Investigator and NTU Associate Professor Kimberly Kline, also the corresponding author of the paper. "The system we designed enables us to easily interrogate various stages during the biofilm developmental cycle of E. faecalis. By selectively silencing certain genes in pre-formed, mature biofilms, we can erode the biofilm and force it to disperse."
The scalable CRISPRi system uses high-throughput screens which can allow for rapid identification of gene combinations to be simultaneously targeted for novel and efficient antimicrobial combinatorial therapies.
The idea behind SMART's inducible CRISPRi system was conceived by Professor Kline and SMART AMR Principal Investigator Professor Timothy Lu, while Dr. Afonina developed and delivered the genetic tool.
More information: Irina Afonina et al. Multiplex CRISPRi System Enables the Study of Stage-Specific Biofilm Genetic Requirements in Enterococcus faecalis, mBio (2020). DOI: 10.1128/mBio.01101-20
Journal information: mBio