Scientists model protein behavior of archaeal viruses to crack protein folding mystery
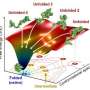
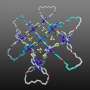
Scientists from the Pacific Quantum Center of Far Eastern Federal University (FEFU) report that the slipknot-structure AFV3-109 protein folds and unfolds depending on temperature. The protein is typical for the archaea, viruses of the oldest single-celled organisms that can survive in the extreme conditions of underwater volcanic sources. The research appears in PLOS ONE.
Using numerical methods and applying quantum field theory that is unique for the study of proteins, the FEFU scientists have probed into the folding topology (scheme) of the AFV3-109 protein featured with a slipknot. The research unfolded in several unexpected and intriguing results.
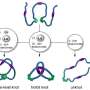
First, it turned out that the sliding knot of the AFV3-109 protein goes through an intermediate knot, which has the topology of a much more complex trefoil knot, a simplest non-trivial knot in mathematics.
Second, before folding of the slipknot is complete, it precedes by the swelling of the almost practically correctly folded AFV3-109 structure in a manner that the free end of the protein can pass into the loop of the knot.
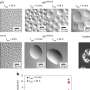
Third, the correct protein structure formation is divided into stages. At the beginning solid secondary structures form, i.e. threads and spirals, and then they fold into a regular knot.
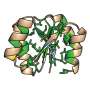
"The knotted structure of proteins makes them more durable and allows viruses, together with archaea, to withstand high temperatures. On the other hand, the presence of a knot makes the protein folding process nontrivial, because the protein cannot fold into the correct three-dimensional structure just by simple random movements of individual parts of the protein backbone. A lot of previous studies carried out by molecular dynamics methods have shown a low probability of such a knot formation, but in nature, this protein always forms a slipknot," says Dr. Alexander Molochkov, head of the Pacific Quantum Center.
For the long AFV3-109 protein molecule that ties itself into a knot, the coordinated collective behavior of the molecule as a whole is necessary. Such behavior turns the protein into an essential model to study mechanisms of folding's complex topology formation. The recent remarkable advances in protein structure prediction by machine learning methods still do not reveal the nature of this structure's formation.
"In our work, we investigate the laws of symmetry that govern the behavior of a protein molecule. We managed to find out that local and chiral symmetry properties completely determine these complex processes and the non-trivial form of the protein," says Alexander Molochkov. "This further confirms that every part of the protein is critical for the entire molecule to function properly. It also means that field theory is relevant for modeling the behavior of proteins that underlie all life."
Following classical field theory, each atom's motion can be considered a part of a collective degree of freedom with a certain number of common coordinates, such as a kink or soliton. An example of such a soliton is a tsunami wave with its destructive power.
Therein, a protein behaves as a whole akin tsunami. Removing a fragment of the protein causes the entire molecule to stop working correctly. The task of scientists is to detect which area to deactivate. That could be the key to understanding the nature of many diseases triggered by protein misbehavior, including cancer, type 2 diabetes, infectious dementia syndrome (in which proteins called prions cause dementia), and enveloped viruses, including the novel coronavirus, Ebola and HIV.
Previously, FEFU researchers modeled the WW-domain's behavior of the FBP28 protein and found out how the replacement of individual amino acids leads to the rearrangement of the protein's entire structure and the consequences of changes in specific amino acids in certain places in the molecular chains.
For the first time, FEFU scientists applied field theory to investigate proteins to predict the environment temperature and acidity-dependent changes in myoglobin structure. The paper explained the release of oxygen molecules when acidity at a certain site of myoglobin changes.
More information: PLOS ONE (2021). DOI: 10.1371/journal.pone.0244547
Journal information: PLoS ONE
Provided by Far Eastern Federal University