Researchers reveal effects of chemical lysis and mechanical lysis on quality of microbial DNA

Yield, purity and integrity, of microbial DNA extracted from digesta samples is crucial for downstream analysis of amplicon sequencing. These markers of quality are influenced by chemical and mechanical lysis. However, contributions of chemical and mechanical lysis have not been investigated in DNA extraction methodology.
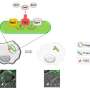
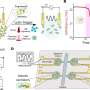
Recently, researchers from the Institute of Subtropical Agriculture (ISA) of the Chinese Academy of Sciences compared the effects of chemical and mechanical lysis on microbial DNA quality and downstream amplicon analysis. The chemical lysis includes QIAamp Fast DNA Stool Mini Kit (QIA) and RBB + C (YM), while the mechanical lysis includes bead (BB) and sand beating (SB).
The study was published in Frontiers in Microbiology.
The researchers found that chemical lysis (QIA and YM) showed similar efficiency to harvest DNA. Although chemical lysis did not affect the overall bacterial community, YM increased some fibrolytic bacterial genera with relative abundance being ranged from 0.33 to 0.78%. The QIA increased the unclassified protozoa than YM about 24-fold and tended to generate more protozoal amplicons and had higher richness in amplicon length polymorphism than YM.
As for mechanical lysis, both BB and SB had similar efficiency to harvest DNA yield and had no difference in DNA quality and bacterial and protozoal community. BB greatly increased DNA yield without affecting DNA quality and richness of bacteria but decreased the richness of protozoa.
These findings revealed that DNA extraction methodology was slightly different for analyzing the bacterial and protozoal community.
Chemical lysis provided by YM and QIA was better to extract DNA for analyzing bacterial and protozoal community, respectively. Sand could be an alternative beater for DNA extraction, and mechanical lysis was not recommended for protozoal community analysis.
More information: Zhi Yuan Ma et al. Effects of Chemical and Mechanical Lysis on Microbial DNA Yield, Integrity, and Downstream Amplicon Sequencing of Rumen Bacteria and Protozoa, Frontiers in Microbiology (2020). DOI: 10.3389/fmicb.2020.581227
Journal information: Frontiers in Microbiology
Provided by Chinese Academy of Sciences