Dynamic risk management in cell populations
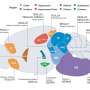
![Dynamic risk management by cell populations. In the absence of environmental signals, uncertainty about the environment is high (middle panel) and the population of E. coli cells were found to demonstrate strong diversity in their sensory response (indicated by broad distribution of response coefficient K_(1/2) in lower panel), enabling responses to many different signals (schematized by color of cell cartoons within top panel in the ‘bet-hedging’ regime). But once a given signal [L] (green curve in top panel) exceeded a threshold (〖[L]〗_0^*) predicted by the mathematical model, the sensory diversity collapsed (indicated by narrow distribution of K_(1/2) in lower panel) , enabling the entire population to ‘focus’ on that particular signal (‘tracking’ regime in top panel). Credit: K. Kamino Dynamic risk management in cell populations](https://scx1.b-cdn.net/csz/news/800a/2020/dynamicriskm.jpg)
Much like investors in the stock market, cell populations prepare for changes in the environment by spreading the risk. The tool box they use contains a repertoire of sensory receptors on the surface of individual cells. These receptors can be tweaked to make individual members of the population responsive to different environmental signals. It was thought that cells could only modify this diversity relatively slowly, by producing new receptor proteins or degrading them. Scientists at AMOLF (Amsterdam, the Netherlands) and Yale University (New Haven, CT) now report the discovery of a mechanism that enables cell populations to tune their diversity much faster, by a combination of physical and chemical interactions between existing proteins. The findings are published in the journal Science Advances on November 13.
The new experimental findings reveal that populations of E. coli bacteria cells maintain a high diversity in their sensory portfolio when environmental signals are absent, but this diversity decreases drastically (by a factor of 10) when subjected to new environmental signals. "This makes a lot of sense," explains Tom Shimizu, a group leader at AMOLF and senior author of the study. "When environmental signals are scarce, uncertainty about the future is at a maximum, so the smart thing to do is to spread your bets broadly across many possible new signals. But the game changes once you sense a certain stimulus. Evidently, cell populations use that new information to 'focus' their attention to a particular signal, so that the whole population can respond together."
What was surprising, however, was how rapidly cells could switch to this new strategy. In biological cells, updates of the portfolio of sensory capabilities were until now assumed to require changes in gene expression, a time consuming process that involves the synthesis of new protein molecules or degradation of old ones. "However, these cell populations could adjust the way they spread their bets over different sensory modes within seconds," says Keita Kamino, lead author of the study who initiated the study at AMOLF and finished it at Yale. "This immediately ruled out the possibility that the cells were updating their bets by gene expression, which takes minutes to hours."
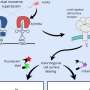
To investigate the mechanism at work, the team then built a mathematical model of the observed diversity, based on known interactions between receptors and other molecules that process and relay signals across the cell. Although the model does not include a way for cells to change their gene expression, they found that it very accurately predicts the changes in the sensory diversity of cells observed in the earlier experiments. "Remarkably, diversity tuning emerges from the a combination of allosteric interactions and covalent modifications between protein molecules," says Thierry Emonet, Professor at Yale University and co-corresponding author of the study, referring to the physical influence of proteins in contact with one another to cause changes in their shape (known as "allosteric interactions"), and the addition and removal of chemical groups to those same receptors (known as "covalent modification"). "Although a model capturing these interactions had already been established by a large body of experimental work, the consequences of this allostery for diversity tuning had not been appreciated."
Thus, the team identified a simple explanation for the fast changes in risk spreading achieved by the cell population. Instead of building and breaking down protein molecules, the mechanism works through chemical modifications of existing proteins and their effects on the physical cross-talk between those proteins, allowing a much faster shift. Such protein modifications are used widely to process and relay signals inside individual cells, but it was not previously known that they could also play a role in the diversification of entire cell populations. Because such modifications are so prevalent throughout cells of all organisms, the authors believe this basic mechanism for fast changes in cellular diversity could be at work in many cell types across biology.
More information: K. Kamino, J.M. Keegstra, J. Long, T. Emonet, T.S. Shimizu, Adaptive tuning of cell sensory diversity without changes in gene expression, Science Advances, advances.sciencemag.org/lookup … .1126/sciadv.abc1087
Journal information: Science Advances
Provided by AMOLF