Researchers use gold nanorod scattering to identify immune system's 'killer and savior'
Every biological system is naturally equipped with a defense mechanism to protect against abnormal changes caused by either local, environmental, or biochemical alteration. White blood cells (WBC) play the role of such a 'soldier' in our immune response. One type of WBC, known as macrophages, is the most efficient and specialized fighter since it is simultaneously equipped with the power of selective identification and elimination of foreign invaders, as well as the potency to repair wounds. Depending on their work distribution, macrophages are mainly comprised of two types, M1 and M2. M1 cells act as the 'professional killer', while M2 cells are more concentrated on healing activity.
In a normal, healthy situation, the immune system maintains a good balance between M1 and M2 cells. But in diseased conditions like bacterial, virus or parasite infections, or inflammations for atherosclerosis, cancer, or arthritis, the balance between M1 and M2 becomes affected, and depending on the crisis, a particular shift in M1 or M2 population occurs. If such changes could be monitored, it would lead to easy diagnostics and prediction of health conditions. There is currently no tool that can provide easy detection of M1/M2 cells directly from tissue fluid or a blood sample in a label-free manner without fluorescent tagging.
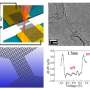
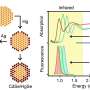
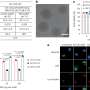
In a study just published in the journal Nano Letters, researchers from Bar-Ilan University in Israel have shown a simple solution to this issue with the help of the scattering effect of Gold Nanorods (GNRs). Gold-based nanoparticles are well known for their prominent optical property with high absorbance and scattering effects. By manipulating the scattering effect and adjusting the surface coating of GNRs, the researchers were able to identify changes in the optical property of M1 and M2 macrophages and utilize them as a parameter to monitor physiological changes.
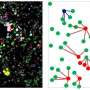
The researchers used the flow cytometer (FCM) to capture changes in the granularity of the cells in order to identify GNR-laden macrophages and determine the specific scattering of GNRs. The FCM is generally used to identify a particular population of fluorescence-labeled cells, but in this case, it was used in label-free detection based only on scattering that came from the GNRs. With this unique method the researchers observed that one type of coating of GNRs exhibited greater selectivity towards M2 cells over M1.
"Our approach in utilizing the scattering of GNRs to identify M1 and M2 macrophages opens a new strategy in cellular identifications using FCM with the help of increased scattering of internalized nanoparticles," says Dr. Ruchira Chakraborty, leading researcher at Prof. Dror Fixler's laboratory at Bar-Ilan University's the Kofkin Faculty of Engineering and Institute of Nanotechnology and Advanced Materials. "Further development of this technique will lead us to build a new point of care or a biopsy tool which can predict the stages of manifestation of diseases like cancer, atherosclerosis, and fibrosis just from the simple tissue fluids or blood samples," says Prof. Dror Fixler, Director of the Bar-Ilan Nano Institute, who led the study in cooperation with Prof. Ran Kornowski and Dr. Dorit Leshem from Beilinson Hospital.
More information: Ruchira Chakraborty et al, The Scattering of Gold Nanorods Combined with Differential Uptake, Paving a New Detection Method for Macrophage Subtypes Using Flow Cytometery, Nano Letters (2020). DOI: 10.1021/acs.nanolett.0c03525
Journal information: Nano Letters
Provided by Bar-Ilan University