Researchers discover first enzymes to catalyze a classic organic reaction
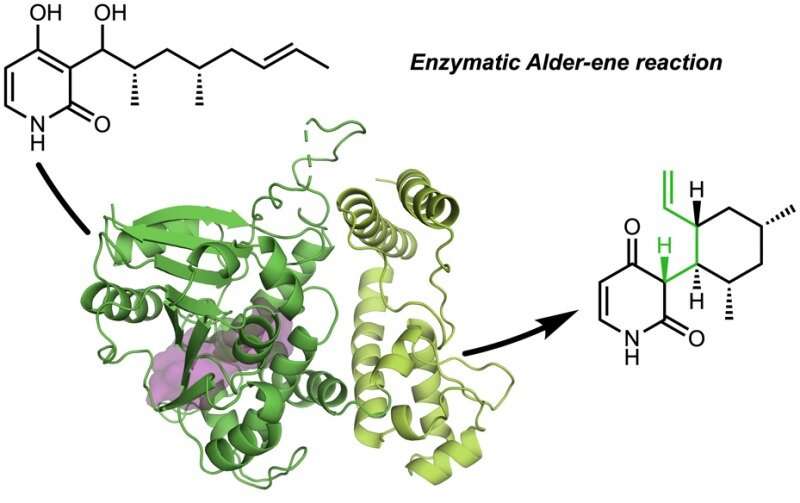
The Tang, Garg, and Houk research groups have discovered nature's natural protein catalysts (enzymes) that catalyze the Alder-ene reaction.
All groups are part of the UCLA Department of Chemistry & Biochemistry. Professor Yi Tang is the Chancellor Professor at the UCLA Department of Chemical and Biomolecular Engineering and also holds a joint appointment in the Department of Bioengineering. Professor Ken Houk holds the Saul Winstein Distinguished Research Chair in Organic Chemistry and Professor Neil Garg holds the Kenneth N. Trueblood Endowed Chair in Chemistry & Biochemistry and serves as the Department Chair.
The paper, "An enzymatic Alder-ene reaction" was published in Nature. The Alder-ene reaction was discovered in 1943, but until now has been used only for chemical synthesis in the laboratory. The Alder-ene reaction is an example of a pericyclic reaction and an analog of the widely-known Diels-Alder reaction. Previously, Tang and Houk identified a new class of enzymes—the pericyclases—for their ability to accelerate pericyclic reactions in primary and secondary metabolism.
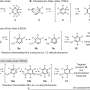
In a collaboration with Jiahai Zhou of the Shanghai Institute of Organic Chemistry, the Tang-Garg-Houk groups reported the structures and mechanisms of two groups of enzymes that catalyze a stereoselective dehydration and concomitant pericyclic reaction, shown below, where the small symbols like PdxI indicate natural enzymes (protein catalysts) discovered by the Tang group. One group of enzymes catalyze an Alder-ene reaction, and the second catalyze hetero-Diels–Alder reactions.
-
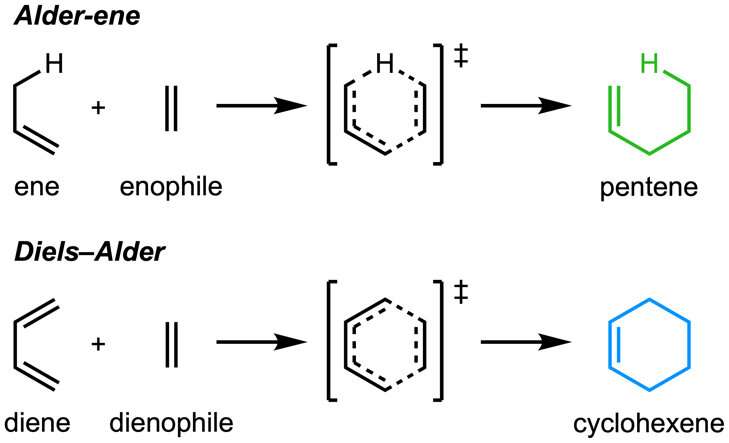
Credit: University of California, Los Angeles -
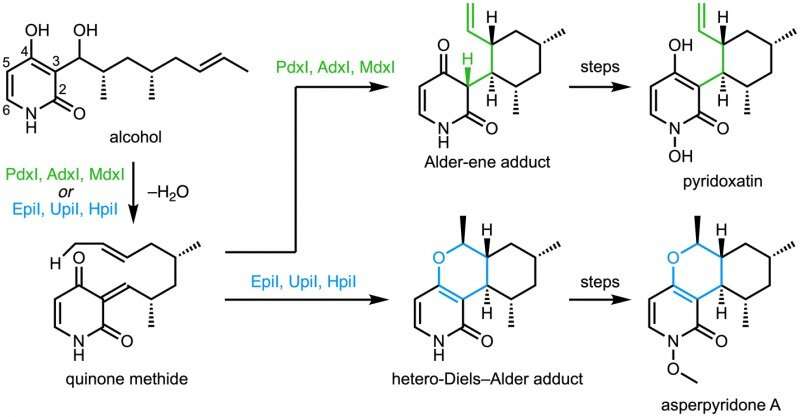
Credit: University of California, Los Angeles
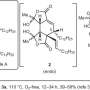
The paper describes the divergent synthesis of pyridoxatin and asperpyridone A. Starting from the alcohol substrate, the enzymes facilitate a dehydration to form a reactive quinone methide intermediate from which the pyridoxatin and asperpyridone A skeletons can form by Alder-ene or Diels-Alder reaction. Masao Ohashi in the Tang group, and Cooper Jamieson in the Houk and Tang groups worked together to identify the factors in the enzymes that control which reaction occurs. Computational predictions guided experimental modifications that altered the enzymes so as to catalyze the other reaction. This type of protein engineering promises to influence laboratory synthesis of related natural products in the future.
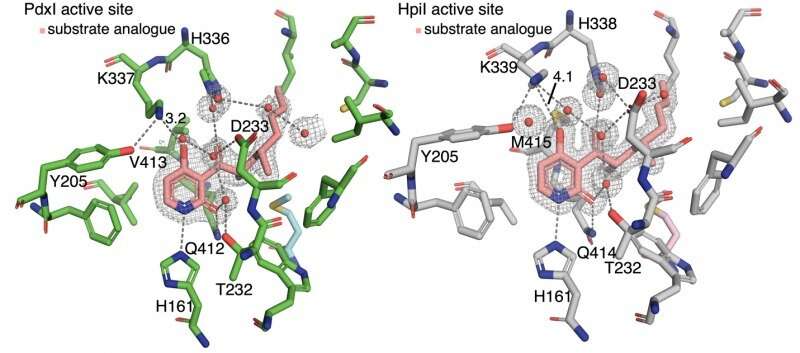
The authors discovered that hydrogen bonding to the pyridone C4 controls reaction type. In enzymes PdxI, AdxI, and MdxI, a lysine residue (K337) hydrogen bonds to C4. This decreases the nucleophilicity of the oxygen and disfavors the hetero-Diels–Alder reaction. PdxI, AdxI, and MdxI use this strategy to selectively form the pyridoxatin skeleton. Enzymes EpiI, UpiI, and HpiI block this hydrogen bond from occurring with a flexible methionine residue (M415) and thereby facilitate the formation of the asperpyridone A skeleton. This subtle geometric difference in the active sites makes a rather large electronic change in the substrate and controls the enzymatic pericyclic reaction.
More information: Masao Ohashi et al. An enzymatic Alder-ene reaction, Nature (2020). DOI: 10.1038/s41586-020-2743-5
Journal information: Nature
Provided by University of California, Los Angeles