Cancer-linked enzyme mechanism newly characterized in study
A new study led by scientists at IUPUI and Indiana University Bloomington is the first to describe a biochemical mechanism that increases the activity of a molecule whose presence is observed in many types of cancer.
The molecule, an enzyme called Pif1helicase, plays a role in many important cellular processes in the body. Tightly regulating this protein is vital to genome stability because too little—or too much—activity can influence aging and age-related diseases, primarily cancer. A common cancer therapy, HDAC inhibitors, can also trigger a spike in this enzyme.
"Currently, we're giving people drugs that increase Pif activity without fully knowing how it affects other parts of the cell that play a role in genome stability," said Lata Balakrishnan, an associate professor of biology in the School of Science at IUPUI, who is co-lead author on the study.
"HDAC inhibitors upregulate certain tumor-suppression genes, and therefore are used in combination therapies to treat specific cancers, but when it comes to their impact on other parts of the cell, we're basically operating in the dark."
The study's other lead author is Matthew Bochman, an associate professor in the IU Bloomington College of Arts and Sciences' Department of Molecular and Cellular Biochemistry. Other co-authors are Christopher Sausen and Onyekachi E. Ononye, Ph.D. students in Bochman's and Balakrishnan's labs, respectively, at the time of the study.
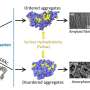
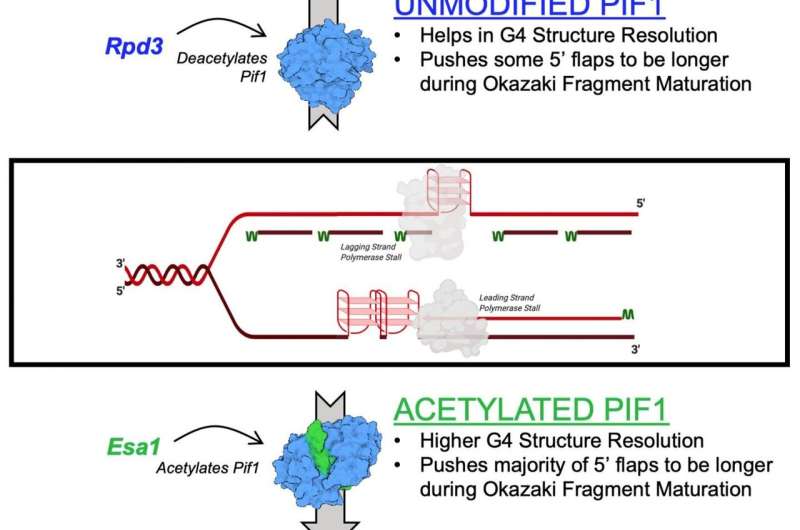
The mechanism described in the study is the effect of lysine acetylation on Pif1. Lysine acetylation occurs when a small molecule called an acetyl group binds to lysine, an amino acid used to build common proteins in the body. This action transforms lysine from a positively charged molecule to a neutrally charged molecule. This neutralization can impact protein function, protein stability and protein-protein interaction in cells, among other things.
Helicases are known as the genetic "zippers" of cells because they pull apart DNA for the purpose of genetic replication and repair. They also help maintain telomeres, the structure at the end of chromosomes that shorten as people age.
In the new study, the researchers identified lysine acetylation on Pif1 helicase and showed the addition of the acetyl group increases the protein's activity—as well as its "unzipping" function. They also found that lysine acetylation changes the shape—or "conformation"—of the Pif1 protein. They believe that this shape change increases the amount of Pif1 helicase.
"The dynamic interplay of the addition and removal of the acetyl group on lysine regulates a wide variety of proteins within the cell," Balakrishnan said. "Perturbations to this process can play a role in cancer, aging, inflammatory responses and even addiction-related behaviors."
"As a class, helicases are involved in a lot of processes necessary for genome integrity," Bochman added. "Any significant failure in these processes is generally carcinogenic."
The precise details of lysine acetylation in Pif1, its effect of the enzyme's shape and the resulting impact on helicase activity took nearly five years to observe and report. The study, carried out in parallel on two IU campuses, was made possible by the lead scientists' complementary expertise. As a biochemist who has previously studied lysine acetylation in other proteins, Balakrishnan was able to isolate Pif1 in vitro to observe its response to chemical reactions in a test tube. In contrast, as a geneticist working in yeast as a model organism to study Pif1, Bochman was able to modify cells in vivo to watch reactions play out in a living organism.
"The ability to observe these reactions in a living cell is often more relevant, but it's also a lot messier," Balakrishnan said. "Our experiments were constantly informing each other as to where to go next."
Looking to the future, Bochman said intricate knowledge of cellular processes—such as lysine acetylation—will increasingly play a role in personalized therapy.
"If you sequence a patient's tumor, you can fine-tune drugs to target very specific enzymes," he said. "Instead of a drug that broadly affects the whole cell, it will be possible to take a targeted approach that reduces potential side effects. This level of personalization is really the future of cancer biology and cancer medicine."
More information: Onyekachi E Ononye et al, Lysine Acetylation Regulates the Activity of Nuclear Pif1, Journal of Biological Chemistry (2020). DOI: 10.1074/jbc.RA120.015164
Journal information: Journal of Biological Chemistry
Provided by Indiana University