Study reveals DNA 'grammar'

DNA three-dimensional structure is determined by a series of spatial rules based on particular protein sequences and their order. This was the finding of a study recently published in Genome Biology by Luca Nanni, Ph.D. student in Computer Science and Engineering at Politecnico di Milano, together with Professors Stefano Ceri of the same University and Colin Logie of the University of Nijmegen.
The first author of the study, Luca Nanni said, "Our study's greatest innovation lies in having identified precise rules for the disposition of CTCF proteins. The beauty and simplicity of CTCF's grammar shows us how nature and evolution produce regularity and incredibly ingenious and functional systems." "Knowing these rules allow CTCF sequences to be engineered to obtain the desired DNA three-dimensional structure. For example, it should be possible to make two disconnected genes interact. Molding DNA structure will open doors to the creation of pharmaceuticals for the treatment of diseases such as cancer."
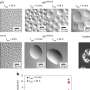
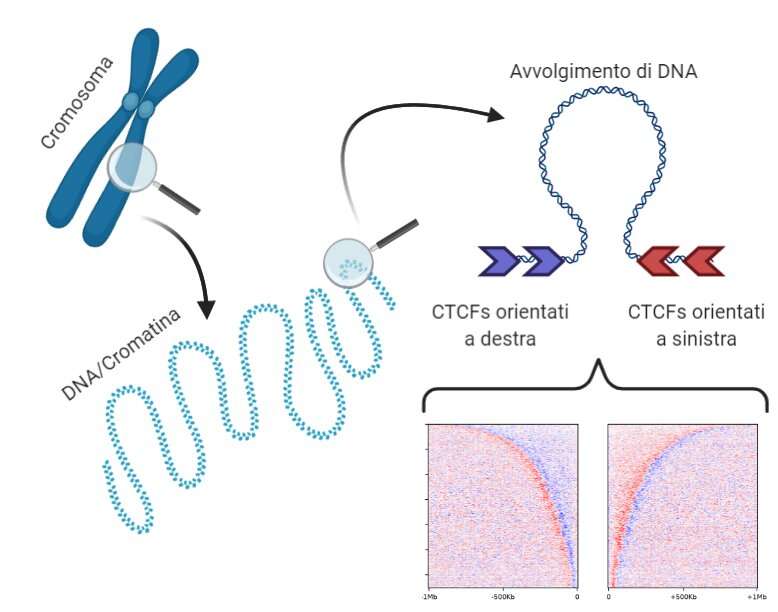
The DNA molecule, which would be about two meters long if completely unrolled, wraps itself based on a complex system that maintains its accessibility and correct reading to reside in the cell's nucleus. Crucial in the study of the three-dimensional structure of the genome are topological domains, which are thought to aggregate DNA zones with similar roles and behavior. For example, genes with similar function are likely to reside in the same topological domain. Nanni continued: "We focused on some specific DNA sequences that encode for the CTCF protein." "This protein isolates portions of DNA creating barriers between the various topological domains. With the help of computer simulations and the creation of a model for classifying these proteins according to their orientation, we identified a surprising regularity in their arrangement along the DNA sequence." The study showed that the orientation and order of these DNA sequences makes it possible to reconstruct topological domains. The human genome compresses following a 'grammar' logic comprising CTCF sequences, orientation, and the distance between them.
More information: Luca Nanni et al, Spatial patterns of CTCF sites define the anatomy of TADs and their boundaries, Genome Biology (2020). DOI: 10.1186/s13059-020-02108-x
Journal information: Genome Biology
Provided by Politecnico di Milano