European and American maize: Same same, but different
German researchers decoded the European maize genome. In comparison to North American maize lines, they discovered variations that underlie phenotypic differences and may also contribute to the heterosis effect. A better understanding of the effect could impact breeding for higher yields. For cultivation of maize in areas with low yields and for challenges imposed by the climate change these observations may be of particular importance.
The maize genome tells an intriguing story about domestication and the shaping of the genome by human selection. Around 10,000 years ago, Native Americans started to domesticate maize in what is Mexico today. They created the basis for one of today's most important sources of food for both humans and livestock. After the discovery of the 'new world' by Columbus, maize was brought from the Americas to Europe. Maize adapted to new growing and climate regimes through directed breeding and selection and finally spread around the globe.
Due to its history, today's maize lines do not only differ in appearance, their genome contains many differences (presence and absence of genes as well as structural variations). In 2009, researchers decoded the genome of the North American maize accession B73. This reference sequence, however, only covers a small part of the global maize genome (pan-genome) and is of limited use as a benchmark for European lines. In order to improve maize breeding and adapt to climate change, basic research on the genome of other maize lines is needed.
European maize genome decoded for the first time
German researchers now succeeded in decoding the European maize genome. They analyzed four different European maize lines using modern sequencing technologies and bioinformatics approaches. In comparison with two lines from North America, they found significant differences in the genetic content and genome structure of these lines—after a few hundred to a thousand years of genetic separation only.
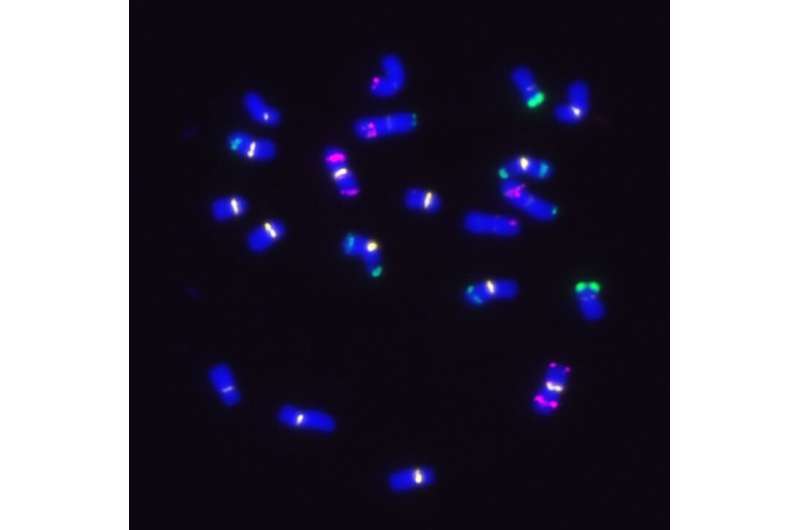
Moreover, so-called 'knob' regions (condensed chromatin regions in the maize DNA) vary substantially in those maize lines. Knob regions are known to affect adjacent genes. In areas where knobs tend to be more pronounced, surrounding genes cannot be read. This results in a loss of genetic function.
Potential cause for heterosis
"We hypothesize that differences in gene content, gene regulation and the influence of knob regions might cause the heterosis effect," says Prof. Klaus Mayer, genomicist at Helmholtz Zentrum München and honorary professor of TUM School of Life Sciences at the Technical University of Munich.
Heterosis occurs when the descendants of crossbreeds are significantly larger and produce higher yields than their parents. If specific genes of a parental generation, e.g. those which determine the height of the maize plant, are not present in a certain region or cannot be read, this will affect the height of the offspring as well. Through crossbreeding with a plant that contains the necessary genetic factor, the defect can be compensated in the next generation. "This results in larger plants with higher yields—without the parents showing these characteristics. In some crossings, this effect can even result in doubling the yield. Although it has been exploited in breeding for a long time, the genetic and molecular basis of heterosis is not yet fully understood," says Prof. Chris-Carolin Schön, professor of Plant Breeding at TUM.
"In a next step, we will test our hypothesis. To this end, we will not only analyze the genomes of the different maize lines, but focus on potential epigenetic processes that may affect the functionality of particular genes," adds Klaus Mayer.
If the researchers' hypothesis proves right, heterosis could be applied even more effectively in future maize breeding. Areas with low yields could benefit from heterosis. Furthermore, these findings could become highly relevant in view of a growing world population and climate change, which poses increasing challenges onto agricultural production.
The study is published in Nature Genetics.
More information: European maize genomes highlight intraspecies variation in repeat and gene content, Nature Genetics (2020). DOI: 10.1038/s41588-020-0671-9 , www.nature.com/articles/s41588-020-0671-9
Journal information: Nature Genetics
Provided by German Research Center for Environmental Health