Maize population study finds genes affected by long-term artificial selection

Researchers conducted a genome-wide scan of a long-term maize breeding study to find the genes involved in increasing the number of ears per maize plant.
The study demonstrates how significantly reduced costs associated with sequencing and the ability to detect common single nucleotide sequence variations (SNPs) within a population are enabling researchers to identify selected genomic regions targeted by artificial selection in natural populations.
One of the projects associated with the goal of converting plant biomass into biofuel is improving biomass production. Long-term breeding projects have provided agricultural researchers with the resources to identify the genes impacted by artificially selecting for specific characteristics. A collaboration involving researchers from the Great Lakes Bioenergy Research Center and the U.S Department of Energy Joint Genome Institute took advantage of one such long-term breeding study in a maize population to conduct such a search.
As reported in the March 1, 2014 issue of Genetics, the team focused on the Golden Glow maize population, which, over 30 generations, had been bred to increase the number of ears per maize plant more than threefold. As populations undergo selection such as the increase of ears per plant, changes in allele frequency occur. Alleles are alternative forms of a gene occupying a specific spot or locus on a chromosome. Changes in allelic composition can provide researchers with information on the genetic control of a trait.
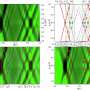
To learn more about these allele frequencies, leaf tissue from maize plants was extracted for SNP genotyping and for whole-genome resequencing. Across the 10 maize chromosomes, 28 "highly divergent" regions were identified, 22 of which contain 5 or fewer annotated gene models, while 14 contain one or zero annotated genes. For most regions, the researchers found that selection appeared to operate on standing genetic variation. For about a quarter of the regions, however, the team found that "selection operated on variants located outside of currently annotated coding regions." This finding, the researchers noted, could either mean the genes aren't present in the reference genome, or these are examples of selection on nongenic DNA.
By combining genomics and bioinformatics approaches in a collaborative setting, researchers hope to improve the efficiency of breeding crop species for biofuel feedstock use, which in turn would contribute to the increased use of renewable energy sources.
More information: Beissinger TM et al. "A genome-wide scan for evidence of selection in a maize population under long-term artificial selection for ear number." Genetics. 2014 Mar;196(3):829-40. DOI: 10.1534/genetics.113.160655
Journal information: Genetics
Provided by US Department of Energy