The amazing travels of small RNAs

In most organisms, small bits of RNA play a key role in gene regulation by silencing gene expression. They do this by targeting and docking onto complementary sequences of gene transcripts (also RNA molecules), which stop the cell machinery from using them to make proteins. This mechanism is called RNA interference (RNAi), and it is critically important in biology.
Remarkably, the RNAi phenomenon is not necessarily confined to single cells; it can also manifest in other tissues and organs far away from the cell of origin. Researchers have been able to observe it mostly in plants, but also in 'lower' animals such as the nematode worm C. elegans.
Proteins and DNA ruled out
Still, one key question had so far gone unanswered: Which messenger substance traverses cells and tissues? "We were able to rule out proteins 20 years ago, once it was discovered that RNAi can travel in plants," says Olivier Voinnet, Professor of RNA Biology at ETH Zurich. RNAi requires that the messenger docks to a complementary sequence of the gene transcript to be silenced. "Proteins alone don't have this capability. DNA leaving the cell nucleus is also unlikely," Voinnet continues. "The most likely candidate has always been an RNA molecule." What has been unclear until now is which precise type and form of RNA—long, short, single- or double-stranded, bound to proteins or not.
Double-stranded fragments travel far and wide
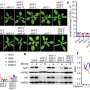
But now, the ETH researchers are shedding light on this process in a new study. They are the first to demonstrate unequivocally that these distant messengers in plants are short double-stranded RNA molecules. These consist of pairs (or double-strands) of just 21 to 24 nucleotides (the building blocks of RNA) called small interfering RNAs, or siRNAs for short. The team's paper was recently published in the journal Nature Plants.
siRNAs usually emerge as large and complex populations from the genomes of viruses that have infected a cell. But a cell's own genes can also serve as blueprints for these molecules. As a result, cells can use RNAi to silence not only invading viruses but also their own genes.
Because RNAi moves, plants have the amazing capacity to modulate gene expression at a distance. This might be particularly important for them to constantly adapt their new growth, enabling what is called "phenotypic plasticity".
To move or not to move
In their new study, the researchers ruled out the possibility that other types of nucleic acids or complexes composed of RNA and proteins move across plant cells. "We can definitively show that double-stranded siRNAs are necessary and sufficient to induce RNAi in distant cells and tissues of plants," Voinnet says.
Not only did the ETH researchers identify the elusive long-distance messengers, they also show, in their study, how siRNAs move and carry out their function. They found that, as long as an siRNA molecule exists as a free double-strand, it is mobile because it cannot bind to a matching RNA transcript. To bind, it first has to be "uploaded" to a specific Argonaute (AGO) effector protein. Only once bound to the correct AGO protein can the siRNA silence the target transcript; the process eventually destroys the fragment itself. The model plant used for the study has ten different AGO proteins, several of which recognize matching siRNA fragments with specific signatures; these signatures are not homogeneous among the large cohorts of mobile siRNAs produced from viruses or the plant's own genes.
AGO proteins determine siRNA movement patterns
Different AGO proteins occur in distinct cells and tissues. The ETH researchers found that as part of the uploading process, matching AGO proteins "consume" a fraction of siRNAs in the cell of origin, but the non-loaded fraction can exit the cell.
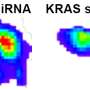
Depending on the presence or absence of certain AGO proteins within the cells traversed by the mobile siRNAs, the molecules, again, will be consumed or not. For example, if there are a plethora of AGO proteins on hand, they will trap plenty of siRNAs with various signatures, essentially stopping movement. If a cell contains hardly any AGO, on the other hand, then most siRNAs will leave and travel greater distances. And finally, if a cell contains large quantities of only one specific AGO, then only those siRNAs with the matching signature will be consumed, while the others will move. In other words, siRNAs are selectively filtered and consumed as they make their way through the plant tissue.
Until now, the plant RNAi community had thought that RNAi moves along linear gradients. However, this does not take into account that AGO proteins selectively use up some siRNAs—but not others—as they move. The new study points out that this consumption process is, in fact, anything but linear.
Countless movement patterns
"The amount and diversity of AGO proteins in traversed cells coupled to the siRNA-intrinsic signatures function together as a kind of molecular sieve, the form of which may differ from cell type to cell type along the siRNA path. Depending on the spatial configuration of this sieve, a wide variety of siRNA movement patterns can be produced," Voinnet explains. He adds, "Even more interestingly, some AGOs can be induced by stress or developmental signals such that the spatial shape of the sieve can change and evolve at any given time".
The countless movement patterns thus lend the mobile RNAi system almost boundless flexibility and versatility in shaping gene expression across distances. Now that they have understood the process, the team of researchers is trying to engineer artificial sieves in plants as a way to control, with high precision, when and where specific siRNAs can move, a method which could have applications in agriculture.
More information: Emanuel A. Devers et al, Movement and differential consumption of short interfering RNA duplexes underlie mobile RNA interference, Nature Plants (2020). DOI: 10.1038/s41477-020-0687-2
Journal information: Nature Plants
Provided by ETH Zurich