New technique offers chemists unprecedented control in drug research
Leiden chemists have developed a new technique with which they can determine the role of kinases—a group of proteins—in a living cell. This technique makes it easier to find new drug targets for diseases such as cancer and rheumatoid arthritis. The team published the findings in the journal Nature Communications.
Potential drug targets
The group of Mario van der Stelt, Professor of Molecular Physiology, studies different types of proteins involved in diseases in the hope of finding new drug targets. For this research, the team looked at kinases: enzymes that switch other proteins on or off. Kinases are involved in a wide variety of processes such as cell division, signaling, and metabolism and play a role in a wide range of diseases such as cancer and inflammatory diseases.
"More than five hundred kinases are known in humans," says first author Tom van der Wel. "But drugs that are currently on the market have only a minor percentage of these kinases as their intended targets. Those other kinases have potential as new targets." However, their role in the body must first be examined. And that's where the problem lies, says the chemical biologist. "They all look very similar and that makes it complicated to determine the role of an individual kinase. Our new method can help with that."
A clever trick
The Leiden team initially looked at one specific kinase, namely tyrosine kinase feline sarcoma oncogene, or FES for short. The researchers devised a clever trick to determine the role of FES.
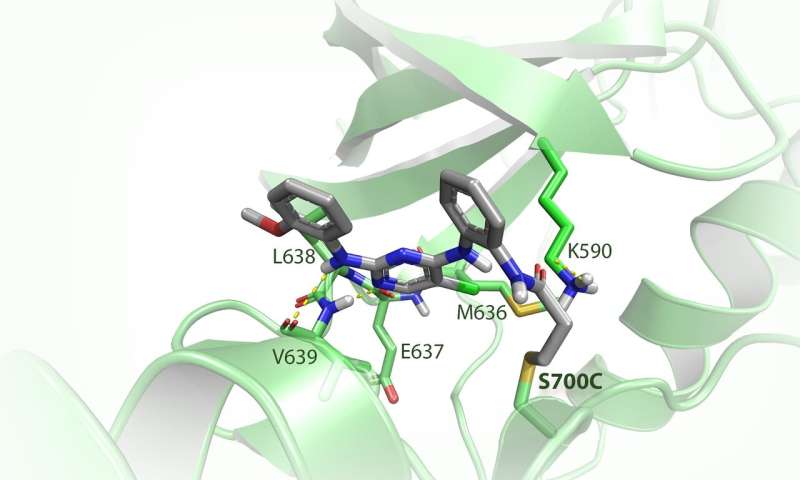
With the gene-editing technique CRISPR-Cas, the researchers changed one of of the 3 billion base pairs in the human genome, resulting in a minuscule adjustment of FES. Van der Wel: "In this way, we changed the amino acid serine into cysteine at the active site of the protein—the place where reactions take place. This effectively amounts to the replacement of one oxygen atom by one sulfur atom out of a total of more than 13,000 atoms in the protein." The newly introduced sulfur atom is more reactive and thus served as a kind of hook to which the researchers could attach their chemical tools.
Van der Wel then designed a fluorescent molecule that reacted specifically with the hook and not with the original serine amino acid. This fluorescent component then served as a sort of searchlight, allowing the chemists to see exactly what FES does in a living cell. "We demonstrated that in the case of an infection, FES is involved in the 'eating' of bacteria by cells of the immune system," says Van der Wel. "Moreover, we discovered that FES activates a specific process in these cells, which also plays a role in cancer and inflammatory diseases such as rheumatoid arthritis. Follow-up research should show whether the inactivation of FES can be used as a new therapeutic approach for these diseases."
Van der Stelt emphasizes that the technique can also be applied to other kinases. "This makes the technique important for validating new drug targets for various diseases." Van der Wel explains why: "We successfully applied the same trick to several other kinases, including FER—a kinase very similar to FES. It took me a long time to develop this method, over four years. But now other scientists can also apply this technique and hopefully contribute to the development of new medicines."
More information: Tom van der Wel et al. Chemical genetics strategy to profile kinase target engagement reveals role of FES in neutrophil phagocytosis, Nature Communications (2020). DOI: 10.1038/s41467-020-17027-5
Journal information: Nature Communications
Provided by Leiden University