Researchers discover intricate process of DNA repair in genome stability

An elaborate system of filaments, liquid droplet dynamics and protein connectors enables the repair of some damaged DNA in the nuclei of cells, researchers at the University of Toronto have found. The findings further challenge the belief that broken DNA floats aimlessly—and highlight the value of cross-disciplinary research in biology and physics.
DNA repair helps ensure genome stability, which in turn allows cells to function and promotes health in all organisms. Double-strand DNA breaks are especially toxic to cells, and researchers had assumed for decades that these breaks floated inside cell nuclei without direction, until they trigger other cellular changes or happen on a fixer mechanism.
That thinking began to change in 2015, when Karim Mekhail and his lab showed that damaged DNA can be intentionally transported by motor protein 'ambulances' to DNA 'hospitals,' areas enriched with certain repair factors in the nuclei. The researchers later worked with U of T aerospace engineers to show that after a single double-strand break, DNA travels for repair via long 'autobahns' of thread-like microtubules, which are also moving.
In the current study, Mekhail and lead author Roxanne Oshidari looked at yeast cells with many DNA double-strand breaks, and showed that coordination between shorter types of microtubule filaments and liquid-like droplets composed of DNA repair proteins enables the creation and function of a DNA repair centre.
"The liquid droplets work with intranuclear microtubules to promote the clustering of damaged DNA sites," says Mekhail, an associate professor of laboratory medicine and pathobiology at U of T. "Repair proteins at these different sites assemble in droplets that fuse into a larger repair-centre droplet, through the action of the shorter nuclear microtubules."
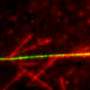
This larger oil-like droplet then behaves like a spider, says Mekhail, shooting out a web of star-shaped filaments that tether to the longer autobahns along which damaged DNA can be transported to the DNA hospitals.
The journal Nature Communications published the findings today.
Mekhail turned to Nasser Ashgriz, a professor in U of T's department of mechanical and industrial engineering, to measure and understand the role of droplets in the repair process. "You couldn't ask for better expertise in fluid dynamics, and he was just across the road," Mekhail says of Ashgriz, who runs U of T's multi-phase flow and spray systems lab.
Mekhail brought a video of the droplets to Ashgriz, who projected it on a large screen in his office and confirmed that fluid dynamics appeared to be at play. But communication across the biology-physics divide was challenging. "Understanding what they do was very difficult in the beginning because our terminologies are totally different," says Ashgriz.
When he and Mekhail used plain language to describe how the droplets behaved, however, things started to make sense. "We focused on the physical aspects of the droplets," Ashgriz says. "The physics that cause their motion and dynamics became our common language."
After months of talks and experiments, computer simulations repeatedly predicted that the shorter filaments would move like pistons, lowering pressure in the nucleoplasm and creating a suction effect that leads to the fusion of droplets. Mekhail and his team confirmed that finding in their lab.
"Often when we dive deep in the specifics of a field, we get separated from one another," Ashgriz says. "Bringing together people with different views can really improve understanding, and this work was a good example—with credit to Karim for his vision and initiative."
Mekhail and his team also uncovered further important properties of the repair droplets with U of T professors Hyun Kate Lee and Haley Wyatt in the department of biochemistry, in a process Mekhail likens to play with toys. They ran the droplets through many tests, bouncing them against each other and observing their behaviour, which turned out to be very similar in a petri dish and in cells.
The most surprising finding came after several cycles of droplet fusion, the researchers found. "It was very bizarre and totally unexpected, I still remember the day," Mekhail says. Oshidari observed that the larger droplets initiate an internal concentration of filament building blocks, forcing creation of a kind of self-interlocking brick road, which together with the spidery webs allow DNA to hook onto the longer autobahn filaments.
The complex process is easy to miss when looking at DNA damage sites, says Mekhail, largely because imaging in the field has become highly automated. Most software has been set up to see what has already been seen. "We can't rely on the old ways of observing," he says. "We need to update our software and also go back to looking with the human eye, guided by simulations when needed."
More information: Roxanne Oshidari et al, DNA repair by Rad52 liquid droplets, Nature Communications (2020). DOI: 10.1038/s41467-020-14546-z
Journal information: Nature Communications
Provided by University of Toronto