Innovative method delivers new insights into the stem cell microenvironment
Researchers from the European Molecular Biology Laboratory (EMBL) and the German Cancer Research Center (DKFZ) in Heidelberg, Germany, have developed new methods to reveal the three-dimensional organization of bone marrow at a single-cell level. Since bone marrow harbors blood stem cells responsible for lifelong blood generation, these results and the new method provide a novel scientific basis to study blood cancer. The results have been published in Nature Cell Biology.
In the published study, the group, led by researchers from EMBL and the DKFZ, present new methods that permit them to characterize complex organs. The team focused their research on the bone marrow of mice, as it harbors blood stem cells that are responsible for lifelong blood production. Because of the ability to influence stem cells and to sustain blood production, there is a growing interest in exploiting the bone marrow environment as a target for novel leukemia treatments.
"So far, very little was known about how different cells are spatially organized within the bone marrow and how they interact with each other," explains Chiara Baccin, a post-doc in the Steinmetz Group at EMBL Heidelberg. The new methods permitted the team to uncover the cellular composition, the three-dimensional organization and how the cells interact with each other. In the process, the researchers have identified previously unknown cell types, so-called "niche cells," that are of great importance for the regulation of blood stem cells.
In order to understand which cells can be found in the bone marrow, where they are localized, and how they might impact on stem cells, the researchers combined single-cell and spatial transcriptomics with novel computational methods. By analyzing the RNA content of individual bone marrow cells, the team identified 32 different cell types, including extremely rare and previously unknown cell types. "We believe that these rare 'niche cells' establish unique environments in the bone marrow that are required for stem cell function and production of new blood and immune cells," explains Simon Haas, group leader at the DKFZ and HI-STEM.
Using novel computational methods, the researchers were not only able to determine the organization of the different cell types in the bone marrow in 3-D, but could also predict how cells interact with each other. "It's the first evidence that spatial interactions in a tissue can be deduced computationally on the basis of genomic data," explains Lars Velten, staff scientist in the Steinmetz Group.
"Our dataset is publicly accessible to any laboratory in the world and it could be instrumental in refining in vivo studies," says Lars Steinmetz, group leader at EMBL Heidelberg. The data, which is now already used by different teams all over the world, is accessible in a 3-D-atlas via a user-friendly web app. All gathered information is available for free and will in the future facilitate the refinement of genetic studies.
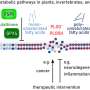
The developed method can be used in different tissues for the study of their 3-D organization. "Our approach is widely applicable and can be used to study the complex pathology of human diseases. We are actually currently in the process of using these approaches on hematological diseases, such as leukemia," highlights Andreas Trumpp, managing director of HI-STEM and division head at DKFZ.
More information: Combined single-cell and spatial transcriptomics reveal the molecular, cellular and spatial bone marrow niche organization, Nature Cell Biology (2019). DOI: 10.1038/s41556-019-0439-6 , nature.com/articles/s41556-019-0439-6
Journal information: Nature Cell Biology
Provided by European Molecular Biology Laboratory