Novel mathematical framework provides a deeper understanding of how drugs interact

Combining two or more drugs can be an effective treatment of diverse diseases, such as cancer. Yet, at the same time, the wrong drug combination can cause major side effects. Currently there is no systematic understanding of how different drugs influence each other. Thus, elucidating how two given drugs interact, and whether they have a beneficial effect, would mean a major step towards drug development to treat diseases more effectively in the future.
On a molecular level, drugs cause complex perturbations of various cellular processes in our body. These processes are orchestrated by an intricate network of molecular interactions, the so-called interactome. Over the last decade, numerous studies have revealed a close relationship between the structure of the interactome and the functional organization of the molecular machinery within the cell. This has opened exciting opportunities for using network-based approaches to investigate the foundations of both healthy and disease states. Following this trend, Principal Investigator Jörg Menche and his group at CeMM developed a novel mathematical framework for mapping out precisely how different perturbations of the interactome influence each other.
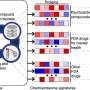
The new study performed by Caldera et al., represents the first general approach to quantifying with precision how drugs interact with each other, based on a mathematical model that considers their high-dimensional effects. Their research reveals that the position of targets of a given drug on the interactome is not random but rather localized within so-called drug modules. They found that the location of a drug module is linked to the specific cell morphological changes induced by the respective treatments, making morphology screens a valuable resource for the investigation of drug interactions. Further they identified various factors that contribute to the emergence of such interactions. Most notably, the distance between two drug modules on the interactome plays a key role: Certain types of interactions are more likely depending on the exact proximity between the two respective drug modules. If the modules are too far away from each other, it is rather unlikely that an interaction will take place.
"We developed a completely new methodology to classify drug interactions. Previous methods could characterize interactions only as synergistic or antagonistic. Our methodology is able to distinguish 12 distinct interaction types and also reveals the direction of an interaction," says Michael Caldera, first author of the study and Ph.D. student in Jörg Menche's Group.
The study of the Menche group has broadened the understanding of how drugs perturb the human interactome, and what causes drugs to interact. Moreover, the new methodology offers the first comprehensive and complete description of any potential outcome that may arise from combining two perturbations. Finally, this methodology could also be applied to address other key challenges, such as dissecting the combined impact of genetic variations or predicting the effect of a drug on a particular disease phenotype. Their research forms a solid base for understanding and developing more effective drug therapies in the future.
More information: Michael Caldera et al, Mapping the perturbome network of cellular perturbations, Nature Communications (2019). DOI: 10.1038/s41467-019-13058-9
Journal information: Nature Communications
Provided by CeMM Research Center for Molecular Medicine of the Austrian Academy of Sciences