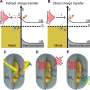
Anchored by a dense neighborhood: What stops cells from going astray
The proteins that constitute the extracellular matrix surrounding a cell exist as fibers. How the spacing between these matrix protein fibers affect clustering of the cell surface receptors integrins and how this influences the formation of integrin-mediated cell-matrix adhesions and subsequent cell spreading was the focus of a recent study led by Dr. Rishita Changede, Senior Research Fellow at the Mechanobiology Institute, National University of Singapore. The study was published in Nature Materials.
Spacing between matrix fibers affects integrin clustering & adhesion formation
Imagine crossing a stream by using rocks that are strewn across it as footholds. Whether you are able to make your way through depends not just on how many rocks there are, but mainly also on how these rocks are positioned along the stream. If your next foothold is even a bit too far away, crossing that stream can become difficult, or sometimes, impossible.
The same applies to cells in our body as they try to attach to surfaces, referred to as the matrix, beneath them. Certain 'receptors' on the outermost layer of the cell, mainly members of the integrin family of proteins, physically interact with partner 'ligand' proteins present in the matrix, such as collagen and fibronectin. The cell basically uses these connections as footholds to spread and move over the matrix or to sense matrix properties. Similar to the rocks in our stream analogy, the 'cellular footholds' need to be optimally spaced in the matrix to promote cell attachment and normal cellular functioning.
Creating various ligand geometries
Within tissues, most ligand proteins are arranged as fibers in the matrix, in varying configurations and densities. The significance of ligand geometry—the specific arrangement of ligand fibers—in promoting the formation and subsequent stabilization of cell-matrix connections was the subject of investigation of a recent study carried out in the Sheetz Lab at the Mechanobiology Institute (MBI), National University of Singapore.
Led by MBI Senior Research Fellow Dr. Rishita Changede and Principal Investigator Prof Michael Sheetz, and involving scientists from Columbia University, U.S., the study employed a technique called electron beam lithography to draw custom, nanosized patterns (made of titanium or gold-palladium lines) on artificial surfaces to mimic ligand geometries found in living tissues.
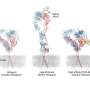
The researchers created either one-dimensional (1-D) or two-dimensional (2-D) nanopatterns. 1-D patterns included single lines, whereas 2-D patterns included paired lines (spaced 50 or 80 nm apart), crossing lines (intersecting at a 25 degree angle), and hexagonal dot patterns (dots spaced 40 nm apart). After patterning, the nano-lines were coated with ligand proteins and the research team microscopically observed and measured how connective tissue cells known as fibroblasts grew on different geometries.
Integrin engagement on 1D and 2-D patterns
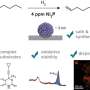
Previous work by Dr. Changede demonstrated that as few as four integrin molecules come together to form clusters that are typically 110 nm in size. These nascent integrin clusters function as basic modules that initiate cell engagement with ligands to form larger cell-matrix connections. Therefore, the researchers theorized that only those nanopatterns in which the ligands are spaced apart by less than 110 nm will allow stable integrin engagement and subsequent cell spreading.
Ligand geometry as the critical factor for cell spreading
Consistent with this, the researchers noted differences in the extent of integrin cluster engagement and cell spreading, based on the ligand geometry on each of these nanopatterns: 1D single lines that were spaced 250 nm or 500 nm apart did not support integrin cluster engagement and cell spreading; however, when the lines were spaced 160 nm apart (slightly more than the integrin cluster size), a few connections were formed and cells were able to spread to a certain extent. On the other hand, 2-D patterns, including paired and crossing lines and hexagonal dots, supported significant integrin cluster engagement and cell spreading.
Notably, such higher integrin cluster engagement and cell spreading occurred on 2-D patterns despite the ligand density (number of ligands in a given area) sometimes being higher on 1D single lines than on 2-D pattern such as hexagonal dots. This observation confirmed a more significant role for ligand geometry over ligand density in controlling the formation of cell-matrix connections and promoting cellular functions such as its spreading and movement along tissues.
Cell-matrix connections are known to be the primary sites for mechanotransduction (the relay of mechanical signals) between a cell and its surroundings; they exert traction forces on the underlying matrix, using them to test the mechanical properties of the matrix. This information is then relayed internally through protein complexes that have been recruited at the connections, in order to effect various changes within a cell.
In the fibrous matrix setting that surrounds cells in a tissue, how these ligand fibers are spaced with respect to each other is of utmost importance in determining how mechanotransduction events are mediated. When the fibers are too close or too far, the integrins are unable to engage stably and initiate the formation of cell-matrix connections. As a result, mechanotransduction pathways go awry, leading to irregular cellular responses that may affect the overall integrity of the tissue. By drawing attention to the importance of ligand geometry in the formation of integrin-dependent connections, the present study adds further details to the molecular mechanisms that govern force mediation through cell-matrix connections, and its impact on cell spreading and movement.
More information: Rishita Changede et al. Integrin nanoclusters can bridge thin matrix fibres to form cell–matrix adhesions, Nature Materials (2019). DOI: 10.1038/s41563-019-0460-y
Journal information: Nature Materials
Provided by National University of Singapore