Tackling E. coli infections
Monash scientists have identified a survival mechanism of bacteria that cause disease in plant and animals, including highly virulent E. coli (Escherichia coli) related diseases.
E. coli are a group of bacteria that are found in the gut of nearly all people and many animals. There are many different strains most are harmless but some can cause serious or lethal infections.
In a study published today in PLoS Genetics researchers show that pathogenic bacteria obtain the essential nutrient iron during infection by pirating it from host proteins.
The study was led by Dr. Rhys Grinter from the Monash School of Biological Sciences, and Professor Trevor Lithgow from the Biomedicine Discovery Institute.
"Our work makes it clear that bacteria have developed sophisticated uptake machineries to obtain the nutrients required for growth," said Dr. Grinter.
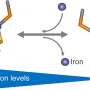
Iron is one such nutrient and is often found inside proteins—the building blocks of hosts cells.
"Bacteria that cause disease in plants can extract iron from plant proteins, by importing the protein and cutting it up once inside the bacterial cell," Dr. Grinter said.
"While it was known that these specific plant infecting bacteria can do this, it was unknown if other bacteria that infect humans and animals are also able to import host proteins," he said.
The research team analyzed the genetic sequences of bacteria and found that genes responsible for importing and processing proteins are widespread in bacteria that cause disease in humans, animals, and plants. They then used cutting-edge techniques to characterize the protein-import systems in molecular detail, confirming their function.
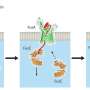
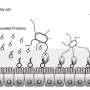
These import systems consist of a channel through the membrane of the bacteria that imports iron-containing proteins and an enzyme inside that bacteria that acts like a pair of molecular scissors. These scissors cut up the target protein once it enters the bacteria releasing the iron it contains so it can be used by the bacteria.
The authors show that the protein-import systems identified in this study are related to that of the virulent plant-pathogen Pectobacterium, which targets iron-containing ferredoxin during infection. However, the study suggests these systems have evolved to target specific proteins present in the host of the bacteria they infect.
"Before this study it was unclear if the ability to import host proteins was widespread or if only Pectobacterium was capable of getting iron in this fashion," Dr. Grinter said.
"We were surprised by how diverse and widespread these systems for protein uptake are."
"Our work suggests they are playing an important role in infection in many pathogenic bacteria."
The research team used X-ray crystallography and the Australian Synchrotron to determine the molecular structure of the purified components of the protein-import system from pathogenic E. coli.
They found major similarities between this system and the ferredoxin import system from Pectobacterium.
"The similarities between these systems in these two very different bacteria were striking," Dr. Grinter said.
"Despite the bacteria adopting radically different lifestyles, separated by millions of years of evolution, it was clear that both systems function in the same way."
The protein import system from E. coli characterized in this study is an important factor in infections caused by pathogenic strains of this bacteria.
"Understanding how this system functions will allow researchers to target and block it to combat these kinds of infections," Dr. Grinter said.
More information: Rhys Grinter et al. Protease-associated import systems are widespread in Gram-negative bacteria, PLOS Genetics (2019). DOI: 10.1371/journal.pgen.1008435
Journal information: PLoS Genetics
Provided by Monash University