Understanding how human cell types develop, vary between individuals, and fail in disease

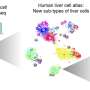
We are in the midst of a fascinating journey to understand the cellular phenotypes that compose human bodies and how the human genome is used to build and maintain each cell. "To catalog our human cell types, to understand how they develop, how they vary between individuals, and how they fail in disease can revolutionize the effort to understand human cell phenotypes," says Gray Camp, Head of the Human Retina and Organoid Development Group at IOB, and co-author of a review in Science.
"Phenotype" can mean many things, but in general, it is a way to classify a set of properties that arise from the interaction of an individual's genotype with its environment.
Why it matters for restoring vision
"Single-cell genomics, as we practice it at IOB in our endeavor to restore vision, is one of the recent advances to develop personalized phenotyping strategies. Single-cell resolved human organ phenotypic maps can be used to identify cell states that are likely most affected by human disease. This can help us overcome one of the major challenges in current drug discovery and development: looking at the population average may mask the crucial and rare disease-causing reaction of a single cell. It may thus hinder the development of accurate disease models, or of detecting patient responses to novel therapies. There are still many obstacles for bringing single-cell phenotyping directly to patients in a clinical setting, but at IOB molecular and clinical research teams closely collaborate daily to overcome this obstacle as soon as possible to restore vision," Gray Camp explains.
Hope to walk into tissue and detect single disease-causing cells
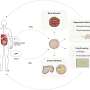
The Science review he wrote in collaboration with Randall Platt and Barbara Treutlein from the Department of Biosystems Science and Engineering, ETH Zürich, in Basel, provides an overview of the current state of single-cell genomic-based phenotyping of human organs and organoids and how CRISPR-Cas technologies will enable phenotypes to be functionally linked to regions of the human genome. The review authors envision that the descriptive phase of single-cell genomics, in which cell phenotypes are cataloged for each healthy human tissue, will culminate in 4-D resolved in silico simulations that enable researchers to walk into the tissue, point to a cell at a location within the tissue at a particular time point, and know its molecular features and its interaction with other cells within the micro-environment.
More information: J. Gray Camp et al. Mapping human cell phenotypes to genotypes with single-cell genomics, Science (2019). DOI: 10.1126/science.aax6648
Journal information: Science
Provided by Institute of Molecular and Clinical Ophthalmology Basel